Abstract
Gene segments from avian H1N1 influenza A viruses have reassorted with other influenza viruses to generate pandemic strains over the past century. Nevertheless, little effort has been invested in understanding the characteristics of avian H1N1 influenza viruses. Here, we present the genome sequence and a molecular and virological characterization of an avian influenza A virus, A/wild bird/Korea/SK14/2014 (A/SK14, H1N1), isolated from migratory birds in South Korea during the winter season of 2014-2015. Full-genome sequencing and phylogenetic analysis revealed that the virus belongs to the Eurasian avian lineage. Although it retained avian-receptor binding preference, A/SK14 virus also exhibited detectable human-like receptor binding and was able to replicate in differentiated primary normal human bronchial epithelial cells. In animal models, A/SK14 virus was moderately pathogenic in mice, and virus was detected in nasal washes from inoculated guinea pigs, but not in direct-contact guinea pigs. Although A/SK14 showed moderate pathogenicity and no evidence of transmission in a mammalian model, our results suggest that the dual receptor specificity of A/SK14-like virus might allow for a more rapid adaptation to mammals, emphasizing the importance of further continuous surveillance and risk-assessment activities.
Similar content being viewed by others
Introduction
Human influenza pandemics occur when an antigenic variant virus with high mammalian transmissibility is introduced into the human population. In the pandemics of the last century, such antigenic shifts have happened through reassortment of gene segments from influenza viruses of different species, including humans, pigs, and birds [32]. Subsequent evolution and adaptation of these variant viruses in the new host species have consequently established new stable lineages of influenza viruses in that particular species. For example, after the 1918 pandemic, influenza viruses diverged into classical swine influenza and seasonal H1N1 viruses in pig and human populations, respectively. The human H1N1 virus remained the main subtype of seasonal human influenza A viruses (IAVs) until the mid-1950s [26]. In 1977, H1N1 IAVs, which were closely related to the H1N1 virus circulating in the early 1950s, reappeared in humans and caused the 1977 Russian pandemic [21]. After 1968, when H3N2 was introduced into the human population, H1N1 IAVs co-circulated with H3N2 IAVs, and both subtypes have since been the main causative agents of seasonal influenza outbreaks worldwide [2, 27]. In 2009, pandemic H1N1 IAVs were first identified in North America and rapidly spread around the world [23]. The 2009 pandemic IAVs were composed of segments from avian, swine, and human IAVs, which were ultimately avian-origin IAVs [29]. In particular, the pandemics in 1918 and 2009 resulted from the direct adaptation of the virus to mammals or the acquisition of several segments from avian H1N1 viruses prior to mammalian adaptation [8, 24]. Recently, Kocer et al. reported that the North American lineage of avian H1N1 influenza viruses contains viruses with high pathogenicity in mammalian hosts [13, 14], suggesting that these avian-origin H1N1 viruses may have a higher propensity to adapt and replicate in mammalian hosts. In this study, we investigated the impact and virological properties of an avian H1N1 influenza virus, A/SK14, isolated from migratory birds in South Korea during the winter of 2014-2015, in mammals. We assessed its pathogenesis, including tissue dissemination, pathology, and virus shedding, in comparison to human H1N1 influenza virus.
Materials and methods
Viruses
A/SK14 virus was isolated from feces of migratory wild birds in Chungnam province of South Korea in the 2014-2015 winter season (GenBank accession numbers KX066868 to KX066875) [16]. To examine the virological and pathogenic properties of A/SK14 virus, we generated a recombinant pandemic H1N1 influenza A/California/04/09 virus (A/CA/04), a well-known and described human H1N1 virus, as a positive control, using a reverse genetic system. Eight gene segments of A/CA/04 virus were cloned in the pHW2000 vector, and the constructs were used to transfect co-cultured 293T and Madin-Darby canine kidney (MDCK) cells. Culture supernatant was collected and used to inoculate 9- to 11-day-old SPF chicken eggs. After virus rescue, the full genome sequences of the rescued viruses were verified by sequencing, and all rescued viruses were stored at -80 °C until use [20].
Cells
MDCK (CRL-2936) and human embryonic kidney (293T, CRL-3216) cells were obtained from the American Type Culture Collection (ATCC) and were grown and maintained in minimum essential medium (MEM) and Opti-MEM (both from Thermo Fisher Scientific, MA, USA) supplemented with 5% fetal bovine serum (Thermo Fisher Scientific, MA, USA), l-glutamine, penicillin (100 U/ml) and streptomycin (100 μg/ml) (Thermo Fisher Scientific, MA, USA). Differentiated primary normal human bronchial epithelial (NHBE) cells were kindly provided by Min-Suk Song and grown in bronchial epithelial cell growth medium (ScienCell Research Laboratories, Inc., CA, USA) according to the manufacturer’s recommendations. All cells were incubated at 37°C in 5% CO2. The 50% tissue culture infective dose (TCID50) was determined by the method described by Reed and Muench [25].
Molecular characterization
Viral RNA was extracted from allantoic fluid from eggs positive for hemagglutination activity using a QIAamp Viral RNA Mini Kit (QIAGEN Inc., Valencia, CA, USA) according to the manufacturer’s instructions. cDNA was generated using a PrimeScript™ First Strand cDNA Synthesis Kit (QIAGEN Inc., Valencia, CA, USA). PCR was conducted using primers targeting the entire coding regions of IAVs using AccuPower PCR MasterMix (Bioneer, Daejeon, Korea) with influenza universal primers [9]. Nucleotide sequences were obtained by direct sequencing using an ABI3730XL DNA Analyzer (Cosmo Gentech, Daejeon, Korea). The sequences were analyzed, compiled using the BioEdit program, and assembled in CLC Sequence Viewer 6.7. Multiple alignments of full coding nucleotide sequences were built with the MUSCLE algorithm [4]. A phylogenetic tree based on the nucleotide sequences was constructed by the neighbor-joining method with 1000 replicates in Molecular Evolutionary Genetics Analysis 6, version 6.06. Reference virus sequences were obtained from the Influenza Virus Resource of the National Center for Biotechnology Information (NCBI).
Studies in mice
For evaluation of the pathogenicity in a mammalian host, six-week-old female C57BL/6J mice (Koatech, Pyeongtaek, South Korea) were inoculated intranasally with 106 EID50/ml of A/SK14 or A/CA/04 virus. To determine the 50% mouse lethal dose (MLD50) of each virus, mice (n = 10) were anesthetized with Avertin (Sigma, MO, USA) and inoculated intranasally with 101 to 108 EID50 in a 30-μl volume. After inoculation, the mice were checked daily for body weight loss for 2 weeks. To assess the pathogenicity of each virus, mice (n = 10) were anesthetized with Avertin (Sigma, MO, USA) and inoculated intranasally with 106 EID50 in a 30-μl volume. After inoculation, the mice were checked for body weight loss daily for 2 weeks. The mice were euthanized if they lost more than 30% of their initial body weight. The MLD50 was calculated by the method described by Reed and Muench [30]. To evaluate viral tissue tropism, inoculated mice were sacrificed on 3, 5, and 7 days postinfection (dpi), and their brain, spleen, liver, kidneys, heart, and lungs were harvested. The tissues were homogenized, and the homogenate supernatant was titrated in a TCID50 assay for virus detection. The limit of virus detection was 1 log10 TCID50/ml. For histopathological analysis, lung tissue from each virus-inoculated mouse was collected on day 5 after inoculation. Lung tissues were fixed in 10% formalin and embedded in paraffin, and sections (about 4-5 µm in thickness) were stained with hematoxylin and eosin (H&E). All mouse experiments were conducted at the Korea Research Institute of Bioscience and Biotechnology (KRIBB, Daejeon, Korea) and were approved by and conducted in accordance with the guidelines of the Institutional Animal Care and Use Committee (IACUC, approval number KRIBB-AEC-16168) of KRIBB.
Studies in guinea pigs
For the assessment of transmissibility, female Hartley guinea pigs weighing 300 to 350 g (Central Lab. Animal Inc., South Korea) that were serologically negative for seasonal influenza viruses were used. The guinea pigs were anesthetized and inoculated intranasally with 106 EID50 of A/SK14 or A/CA/04 viruses. Guinea pigs (n = 3) were euthanized to collect nasal washes for virus titration in MDCK cells. For direct-contact transmission experiments, guinea pigs housed in a cage placed inside an isolator were inoculated intranasally with 106 EID50 of test virus. Twenty-four hours later, three naive animals were introduced into the same cage. Nasal washes were collected daily, beginning on day 2 post-inoculation (p.i.; 1 day after contact). The presence of virus in the nasal washes was determined by TCID50 assay in MDCK cells. For seroconversion, serum samples were collected from guinea pigs 14 days after inoculation, treated with receptor-destroying enzyme (RDE) at 37°C for overnight, heat-inactivated at 56°C for 30 minutes, and then measured by hemagglutination inhibition (HI) assay with 0.5% chicken red blood cells. The limit of detection is 1:10 for the HI assay. All guinea pig experiments were conducted at Chungbuk National University (Cheongju, Korea) and were approved by and conducted in accordance with the guidelines of the Institutional Animal Care and Use Committee (IACUC, approval number CBNUA-937-16-02) of Chungbuk National University.
Growth kinetics
To evaluate the multistep growth kinetics of A/SK14 and A/CA/04 virus in vitro, MDCK and NHBE cells were infected with the virus at a multiplicity of infection (MOI) of 0.01 plaque-forming units (PFU) per cell. After a 1-hour incubation, the virus inoculum was removed, the cells were washed with PBS, and infection medium containing 1 μg of tosylsulfonyl phenylalanyl chloromethyl ketone (TPCK)-treated trypsin per ml was added. Supernatants were collected at 12, 24, 36, 48, and 72 hours postinfection (hpi). The virus titer of each supernatant was determined by TCID50 assay in MDCK cells.
Receptor-binding assay
A/SK14 was examined for its binding preference for avian- and human-specific virus receptors. A solid-phase direct-binding assay [19] was performed for evaluating the virus-binding capacity of the synthetic biotinylated glycans α2,3 (Neu5Acα2-3Galβ1-3GalNAcα-PAA-biotin, Neu5Acα2-3GalNAcα-PAA-biotin, Neu5Acα2-3Galβ1-4GlcNAcβ-PAA-biotin) and α2,6 (Neu5Aca2-6Galβ1-4Glcβ-PAA-biotin, Neu5Acα 2-6Galβ1-4Glcβ-PAA-biotin, Neu5Gca2-6GalNAca-PAA-biotin) as described previously [33]. In brief, 10 μg of fetuin (Sigma, MO, USA) was added to each well of a 96-well microtiter plate, which was then incubated at 4°C overnight and blocked with 5% bovine serum albumin (BSA) in phosphate-buffered saline (PBS), pH 7.4, at room temperature for 1 hour. After removal of the BSA solution, the plates were washed four times with ice-cold PBS and incubated with 64 HAU of each virus per ml at 4°C overnight. After virus removal, the plate was washed as described above and incubated with 0.1 ml of each of the biotinylated glycans per well at different concentrations at 4°C for 3 hours. After washing three times with ice-cold PBS, horseradish-peroxidase-conjugated streptavidin (1000-fold dilution in PBS) was added to each well, and the plate was incubated at room temperature for 1 hour in the dark. After three washes, 0.05 ml of 3,3′,5,5′-tetramethylbenzidine (TMB) substrate was added to each well, and the plate was incubated at room temperature for 10 min. The reaction was stopped with 0.05 ml of 50 mM HCl, and the optical density was then measured at 450 nm in a Synergy HTX multi-mode microplate reader (BioTek, VT, USA). As a control with a binding preference for the avian receptor, we used a low-pathogenic H5N2 avian influenza virus, A/Aquatic Bird/Korea/CN2/2009 (A/CN2/09) [22].
Statistical analysis
Statistically significant differences in multistep experimental groups were determined using analysis of variance (ANOVA) in Graph Pad Prism version 5.03. A P-value of less than 0.05 was considered statistically significant.
Results
Genetic and biological properties of the A/SK14 virus
A virus from fecal samples of wild birds collected in Chungnam province in 2014 was identified by reverse transcription polymerase chain reaction (RT-PCR) and confirmed by genomic sequencing and nucleotide BLAST analysis. Phylogenetic analysis of all gene segments confirmed that the A/SK14 virus belongs to the Eurasian lineage of avian H1N1 IAVs (Fig. 1, Supplementary Fig. 1). In molecular analysis of the hemagglutinin (HA) protein, the 226Q substitution was identified in the receptor-binding site of A/SK14. This mutation has been associated with increased disease severity in 2009 pandemic H1N1 viruses due to acquisition of receptor specificity against α2,6-SA moieties [34]. We did not find additional amino acid sequence motifs previously shown to correlate with antiviral drug resistance and disease severity such as N66 in PB1-F2 [7]; E627 in PB2 [30]; ESEV (PDZ-binding domain), T42, and T91 in NS [17]; G31 in M [23]; H275 and D294 in NA [1] (Table 1). Next, we assessed the biological features of A/SK14 virus in vitro and in vivo, using the well-known human H1N1 isolate A/CA/04 as a positive control. The A/SK14 virus grew well in SPF eggs and MDCK cells, but it was less infectious in mice than the A/CA/04 virus (7.5 vs. 4.0 MLD50) (Table 2).
Phylogenetic analysis of the HA and NA genes of A/SK14 virus. A phylogenetic tree based on nucleotide sequences was constructed by the neighbor-joining method with 1000 replicates in Molecular Evolutionary Genetics Analysis 6. Reference viruses were obtained from the Influenza Virus Resource of the National Center for Biotechnology Information (NCBI). Viruses examined in this report are indicated by a thick black dot
Replication kinetics and receptor-binding preference of A/SK14
To determine the growth properties of A/SK14 in mammalian cells, we performed multistep viral growth kinetics experiments in MDCK and primary NHBE cells. Because the airway epithelium is the primary site of influenza virus replication in humans, we confirmed the ability of the A/SK14 virus to replicate in NHBE cells [3], which predominantly express α2,6-SA-linked receptors. The A/SK14 virus showed growth kinetics similar to those of the H1N1 virus A/CA/04, and there was no significant difference in their viral titers (Fig. 2a). Interestingly, in NHBE cells, A/SK14 showed delayed kinetics at 12 and 24 hours postinfection (hpi) when compared to A/CA/04 virus (Fig. 2b). The titer of A/SK14 virus was approximately ten times lower than that of A/CA/04 virus (p < 0.01). However, at 36 hpi, there was no significant difference in replication efficiency between these viruses, and at 72 hpi, they reached similar titers.
Growth kinetics of A/SK14 virus in MCDK and NHBE cells. Replication of A/SK14 and A/CA/04 virus was confirmed in MDCK (A) and NHBE (B) cells. Cells were inoculated with virus at an MOI of 0.01, and supernatants from the infected culture were collected at different time points. The titers shown are means ± SD from three independent experiments
We examined the receptor binding preference of A/SK14 virus, because the binding affinity of HA for human receptors is an important factor for successful transmission in mammalian species. To determine the glycan-binding specificity of the A/SK14 virus, we measured direct binding to glycans with α2,3-sialyl linkages (α2,3-SAL) or α2,6-sialyl linkages (α2,6-SAL) using a solid-phage glycan binding assay. Our analysis showed that A/SK14 had substantial binding affinity towards α2,6-SAL, albeit less than that of A/CA/04. Unlike A/CA/04, however, A/SK14 and the avian H5N2 virus A/CN2/09 retained their ability to bind to α2,3-SAL-containing glycans (Fig. 3).
Pathogenicity of A/SK14 virus in C57BL/6J mice
Mice have been widely used as a mammalian model for assessing the pathogenesis of influenza viruses, including avian-origin H1N1 [13], highly pathogenic avian influenza (HPAI) H5N1, and human H1N1 viruses. To evaluate the lethality of A/SK14 in mice, 6-week-old C57BL/6J mice (10 mice per each group) were inoculated intranasally with 106 EID50 of A/SK14 or A/CA/04 virus in 30 µl. In terms of pathogenesis, the mice inoculated with A/CA/04 virus showed signs more-severe of illness, such as body weight loss, reduced activity and ruffled fur, and all of the mice died by 7 to 8 dpi. In contrast, A/SK14-inoculated mice exhibited mild weight loss, and two out of 10 animals succumbed to infection on day 9 post-inoculation. (Fig. 4). Next, we assessed tissue tropism in virus-inoculated mice. A/SK14 was detected in the lungs, but at a significantly lower titer (p < 0.001) than A/CA/04. However, both viruses were undetectable in non-respiratory tissues, including the brain, spleen, liver, kidneys, and heart (Fig. 5a). In addition, the histopathological features of pulmonary lesions in mice infected with A/SK14 and A/CA/04 were compared. Consistent with the results indicating lung tissue tropism, A/CA/04-inoculated mice showed severe pulmonary lesions, such as necrotizing bronchiolitis and bronchiolar epithelial hyperplasia, by H&E staining (Fig. 5b). Pulmonary lesions were also observed in A/SK14-inoculated mice, but we found that A/SK14 induced milder lung pathology than A/CA/04 (Fig. 5b). Taken together, these results indicate that the A/SK14 is moderately pathogenic in a mammalian host.
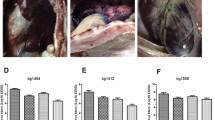
Tissue tropism of A/SK14 virus in C57BL/6J mice. For determination of virus titers in the lungs of infected mice, three mice per group were inoculated intranasally with 106 EID50 of each virus, and lungs were collected on 3, 5, and 7 days post-inoculation (pi). Virus titers were determined by titration in MDCK cells (a). For histopathological analysis, lungs were collected on day 5 pi, fixed in 10% formalin, and embedded in paraffin, and sections were stained with H&E (b). Representative images were taken at a magnification of 200×
Transmissibility of the A/SK14 virus in guinea pigs
As we found that A/SK14 could replicate in NHBE cells and was able to bind to α2,6-linked SAs, we evaluated whether it could be transmitted between mammals of different species using guinea pigs, which have been established as an animal model for such studies [18]. To assess the transmissibility of A/SK14 in mammals, we inoculated guineas pigs intranasally with the A/SK14 and A/CA/04 viruses. As expected, we found that A/CA/04 replicated to high levels up to day 7 in inoculated guinea pigs, with the virus being transmitted to all contact animals (Fig. 6a). While A/SK14 was able to infect the directly inoculated guinea pigs, it did not replicate to the same level as did A/CA/04 and was not transmitted by direct contact (Fig. 6b). We also observed that guinea pigs that were exposed to A/CA/04-inoculated animals demonstrated seroconversion in HI assays performed at 14 dpi (> 80 and > 160 HI units), but we did not detect HI activity in serum from guinea pigs that had been exposed to A/SK14 (Table 2). These results suggested that, despite some α2,6-SAL receptor binding ability, A/SK14 did not have the capacity to be transmitted to mammals.
Transmissibility of A/SK14 virus in guinea pigs. To determine direct-contact transmissibility, guinea pigs were inoculated intranasally with 106 EID50 of A/SK14 or A/CA/04 virus, and naïve guinea pigs (n = 3) were placed in the same cage 24 h later. Nasal washes were collected from each animal, and virus shedding was determined by titration in MDCK cells. Dashed lines indicate the lower limit of detection
Discussion
Asian countries such as China, Vietnam, and South Korea are located along several migratory bird flyways [6]. According to previous reports, the Korean peninsula is prone to outbreaks of avian influenza, including those caused by highly pathogenic H5Nx IAVs [12, 28], low-pathogenic H7Nx [11], and avian H1N1 as well as swine H1N1 [10] IAVs, and pandemic outbreaks of H1N1 IAVs in humans have been repeatedly reported. A recent study found that avian-origin H1N1 IAVs caused high mortality in mammals [13, 15]. Therefore, there is a strong need for biological and molecular characterization of newly emerging H1N1 viruses to evaluate their potential threat to human populations. In this report, we characterized an avian-origin H1N1 virus isolated in Korea in 2014 in terms of its biological and molecular properties.
Genetic analysis indicated that all gene segments of A/SK14 belonged to Eurasian lineages of avian-origin H1N1 viruses. A/SK14 harbored the amino acid sequence PSIQSR/GLF in the HA cleavage site, a characteristic of low-pathogenic influenza viruses. A/SK14 contained avian-type receptor binding signatures at HA positions 190 and 225 (E and G, respectively), but interestingly, the virus contained the mammalian-type receptor signature Q at position 226. The 226Q substitution has been associated with increased pathogenicity and transmissibility of influenza viruses in ferrets [34]. A/SK14 could replicate in primary NHBE cells and reached titers similar to those of A/CA/04 despite delayed kinetics. In addition, A/SK14 also had detectable binding affinity for both avian- and human-specific glycans in a solid-phase direct binding assay. The ability to bind human receptors is likely to be associated with the presence of HA 226Q in A/SK14. Polymerase complex proteins of influenza virus, especially PB2 proteins, have important roles in transmission of avian influenza virus to mammals [31], and the substitutions E627K and D701N are well-defined molecular pathogenic markers associated with increased pathogenicity in mammals [5, 30]. Of the 157 avian H1N1 PB2 sequences (from 2010 to 2016), including A/SK14, in the influenza sequence database (www.fludb.org), none of the viruses contained lysine at position 627 or asparagine at position 701. Previous reports have shown that avian H5N1 influenza viruses isolated from humans had the substitutions 627K and 701N in the PB2 gene [30]. Therefore, we suggested that A/SK14 lacking the PB2 mutation might have limited pathogenicity in mammalian hosts.
We used both mouse and guinea pig models to attempt to assess the zoonotic risk of A/SK14. Unlike A/CA/04, which was transmissibility in guinea pigs, A/SK14 virus was detected in nasal washes from all inoculated guinea pigs but was not detected in the nasal washes from guinea pigs that were in direct contact with infected animals, indicating that no virus transmission had occurred. In our mouse experiments, A/SK14 exhibited moderate pathogenicity compared to A/CA/04. In nature, infection and transmission of AIVs between birds and humans have been limited due to species barriers in the primary site of viral replication. Therefore, further studies of adaptation to mammalian hosts and viral mutagenesis studies are required to better understand the virulence determinants of A/SK14 virus in mammals.
In summary, A/SK14 shows moderate pathogenicity and no transmissibility in guinea pigs. However, it is able to bind to both avian and human receptors and replicate in primary human airway cells. It is thus possible that these mammalian-virus-like properties could enhance the ability of A/SK14-like viruses to adapt more rapidly than other avian influenza viruses to mammalian hosts under selective pressure or subsequent to reassortment events.
References
Baek YH, Song MS, Lee EY, Kim YI, Kim EH, Park SJ, Park KJ, Kwon HI, Pascua PN, Lim GJ, Kim S, Yoon SW, Kim MH, Webby RJ, Choi YK (2015) Profiling and characterization of influenza virus N1 strains potentially resistant to multiple neuraminidase inhibitors. J Virol 89:287–299
Bedford T, Riley S, Barr IG, Broor S, Chadha M, Cox NJ, Daniels RS, Gunasekaran CP, Hurt AC, Kelso A, Klimov A, Lewis NS, Li X, McCauley JW, Odagiri T, Potdar V, Rambaut A, Shu Y, Skepner E, Smith DJ, Suchard MA, Tashiro M, Wang D, Xu X, Lemey P, Russell CA (2015) Global circulation patterns of seasonal influenza viruses vary with antigenic drift. Nature 523:217–220
Chan RW, Yuen KM, Yu WC, Ho CC, Nicholls JM, Peiris JS, Chan MC (2010) Influenza H5N1 and H1N1 virus replication and innate immune responses in bronchial epithelial cells are influenced by the state of differentiation. PLoS One 5:e8713
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucl Acids Res 32:1792–1797
Gabriel G, Dauber B, Wolff T, Planz O, Klenk HD, Stech J (2005) The viral polymerase mediates adaptation of an avian influenza virus to a mammalian host. Proc Natl Acad Sci USA 102:18590–18595
Global Consortium for HN, Related Influenza V (2016) Role for migratory wild birds in the global spread of avian influenza H5N8. Science 354:213–217
Hai R, Schmolke M, Varga ZT, Manicassamy B, Wang TT, Belser JA, Pearce MB, Garcia-Sastre A, Tumpey TM, Palese P (2010) PB1-F2 expression by the 2009 pandemic H1N1 influenza virus has minimal impact on virulence in animal models. J Virol 84:4442–4450
He CQ, Ding NZ, Mou X, Xie ZX, Si HL, Qiu R, Ni S, Zhao H, Lu Y, Yan HY, Gao YX, Chen LL, Shen XH, Cao RN (2012) Identification of three H1N1 influenza virus groups with natural recombinant genes circulating from 1918 to 2009. Virology 427:60–66
Hoffmann E, Stech J, Guan Y, Webster RG, Perez DR (2001) Universal primer set for the full-length amplification of all influenza A viruses. Arch Virol 146:2275–2289
Kang HM, Lee EK, Song BM, Jeong J, Kim HR, Choi EJ, Shin YK, Lee HS, Lee YJ (2014) Genetic and pathogenic characteristics of H1 avian and swine influenza A viruses. J Gen Virol 95:2118–2126
Kang HM, Park HY, Lee KJ, Choi JG, Lee EK, Song BM, Lee HS, Lee YJ (2014) Characterization of H7 influenza A virus in wild and domestic birds in Korea. PLoS One 9:e91887
Kim YI, Pascua PN, Kwon HI, Lim GJ, Kim EH, Yoon SW, Park SJ, Kim SM, Choi EJ, Si YJ, Lee OJ, Shim WS, Kim SW, Mo IP, Bae Y, Lim YT, Sung MH, Kim CJ, Webby RJ, Webster RG, Choi YK (2014) Pathobiological features of a novel, highly pathogenic avian influenza A(H5N8) virus. Emerg Microbes Infect 3:e75
Kocer ZA, Krauss S, Stallknecht DE, Rehg JE, Webster RG (2012) The potential of avian H1N1 influenza A viruses to replicate and cause disease in mammalian models. PLoS One 7:e41609
Kocer ZA, Carter R, Wu G, Zhang J, Webster RG (2015) The genomic contributions of Avian H1N1 influenza A viruses to the evolution of mammalian strains. PLoS One 10:e0133795
Kocer ZA, Krauss S, Zanin M, Danner A, Gulati S, Jones JC, Friedman K, Graham A, Forrest H, Seiler J, Air GM, Webster RG (2015) Possible basis for the emergence of H1N1 viruses with pandemic potential from avian hosts. Emerg Microbes Infect 4:e40
Koo BS, Kim HK, Na W, Song D, Kim DJ, Yoon SW, Jeong DG (2017) Complete genome sequence of an Avian H1N1 influenza virus strain isolated from migratory birds in the Republic of Korea. Genome Announc 5:e00356-17
Liu H, Golebiewski L, Dow EC, Krug RM, Javier RT, Rice AP (2010) The ESEV PDZ-binding motif of the avian influenza A virus NS1 protein protects infected cells from apoptosis by directly targeting scribble. J Virol 84:11164–11174
Lowen AC, Mubareka S, Tumpey TM, Garcia-Sastre A, Palese P (2006) The guinea pig as a transmission model for human influenza viruses. Proc Natl Acad Sci USA 103:9988–9992
Matrosovich MN, Gambaryan AS (2012) Solid-phase assays of receptor-binding specificity. Methods Mol Biol 865:71–94
Na W, Lyoo KS, Yoon SW, Yeom M, Kang B, Moon H, Kim HK, Jeong DG, Kim JK, Song D (2016) Attenuation of the virulence of a recombinant influenza virus expressing the naturally truncated NS gene from an H3N8 equine influenza virus in mice. Vet Res 47:115
Nakajima K, Desselberger U, Palese P (1978) Recent human influenza A (H1N1) viruses are closely related genetically to strains isolated in 1950. Nature 274:334–339
Nam JH, Shim SM, Song EJ, Espano E, Jeong DG, Song D, Kim JK (2017) Rapid virulence shift of an H5N2 avian influenza virus during a single passage in mice. Arch Virol 162:3017–3024
Novel Swine-Origin Influenza AVIT, Dawood FS, Jain S, Finelli L, Shaw MW, Lindstrom S, Garten RJ, Gubareva LV, Xu X, Bridges CB, Uyeki TM (2009) Emergence of a novel swine-origin influenza A (H1N1) virus in humans. N Engl J Med 360:2605–2615
Qi L, Kash JC, Dugan VG, Wang R, Jin G, Cunningham RE, Taubenberger JK (2009) Role of sialic acid binding specificity of the 1918 influenza virus hemagglutinin protein in virulence and pathogenesis for mice. J Virol 83:3754–3761
Reed LJ, Muench H (1938) A simple method of estimating fifty percent endpoints. Am J Hyg 27:493–497
Reid AH, Fanning TG, Hultin JV, Taubenberger JK (1999) Origin and evolution of the 1918 “Spanish” influenza virus hemagglutinin gene. Proc Natl Acad Sci USA 96:1651–1656
Schrauwen EJ, Fouchier RA (2014) Host adaptation and transmission of influenza A viruses in mammals. Emerg Microbes Infect 3:e9
Si YJ, Lee IW, Kim EH, Kim YI, Kwon HI, Park SJ, Nguyen HD, Kim SM, Kwon JJ, Choi WS, Beak YH, Song MS, Kim CJ, Webby RJ, Choi YK (2017) Genetic characterisation of novel, highly pathogenic avian influenza (HPAI) H5N6 viruses isolated in birds, South Korea, November 2016. Euro Surveill 22(1):30434
Smith GJ, Vijaykrishna D, Bahl J, Lycett SJ, Worobey M, Pybus OG, Ma SK, Cheung CL, Raghwani J, Bhatt S, Peiris JS, Guan Y, Rambaut A (2009) Origins and evolutionary genomics of the 2009 swine-origin H1N1 influenza A epidemic. Nature 459:1122–1125
Steel J, Lowen AC, Mubareka S, Palese P (2009) Transmission of influenza virus in a mammalian host is increased by PB2 amino acids 627K or 627E/701N. PLoS Pathog 5:e1000252
Subbarao K, Klimov A, Katz J, Regnery H, Lim W, Hall H, Perdue M, Swayne D, Bender C, Huang J, Hemphill M, Rowe T, Shaw M, Xu X, Fukuda K, Cox N (1998) Characterization of an avian influenza A (H5N1) virus isolated from a child with a fatal respiratory illness. Science 279:393–396
Yoon SW, Webby RJ, Webster RG (2014) Evolution and ecology of influenza A viruses. Curr Top Microbiol Immunol 385:359–375
Yoon SW, Chen N, Ducatez MF, McBride R, Barman S, Fabrizio TP, Webster RG, Haliloglu T, Paulson JC, Russell CJ, Hertz T, Ben-Tal N, Webby RJ (2015) Changes to the dynamic nature of hemagglutinin and the emergence of the 2009 pandemic H1N1 influenza virus. Sci Rep 5:12828
Zhang Y, Zhang Q, Gao Y, He X, Kong H, Jiang Y, Guan Y, Xia X, Shu Y, Kawaoka Y, Bu Z, Chen H (2012) Key molecular factors in hemagglutinin and PB2 contribute to efficient transmission of the 2009 H1N1 pandemic influenza virus. J Virol 86:9666–9674
Acknowledgements
We thank Hyun-Woo Lee and Hai Yen Le for technical assistance.
Funding
This work was supported by grants from the KRIBB Initiative program, supported by the BioNano Health-Guard Research Center, funded by the Ministry of Science, ICT and Future Planning (MSIP) of Korea as Global Frontier Project (Grant no. H-GUARD 2013M3A6B2078954) and supported by Animal Disease Management Technology Development, Ministry of Agriculture, Food and Rural Affairs (Grant no. 316042-03), and supported by Korea Ministry of Environment (MOE) as “Public Technology Program based on Environmental Policy (no. 2016000210002)”.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
All animal experiments were conducted at the Korea Research Institute of Bioscience and Biotechnology (KRIBB, Daejeon, Korea) and Chungbuk National University (Cheongju, Korea) and were approved by and conducted in accordance with the guidelines of the Institutional Animal Care and Use Committee of KRIBB and Chungbuk National University.
Additional information
Handling Editor: Ayato Takada.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Koo, BS., Kim, H.K., Song, D. et al. Virological and pathological characterization of an avian H1N1 influenza A virus. Arch Virol 163, 1153–1162 (2018). https://doi.org/10.1007/s00705-018-3730-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-018-3730-0