Abstract
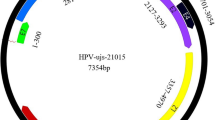
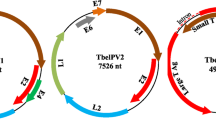
A highly divergent human papillomavirus, named HPV-CH2, was identified in fecal samples from children with diarrhea in China by 454 high-throughput sequencing. Here, we report the complete genome sequence and genetic organization of the virus. The full-length nucleotide sequence of HPV-CH2 shares the highest sequence similarity with HPV-156, with 72 % nucleotide sequence identity. The L1 gene of the HPV-CH2 shared <70 % nucleotide identity with previously reported HPVs, suggesting HPV-CH2 as a new type of papillomavirus. Phylogenetic analysis revealed that the HPV-CH2 belongs to the genus Gammapapillomavirus. No HPV-CH2 was detected by PCR in samples from children with both gastroenteritis and respiratory infection.


Similar content being viewed by others
References
Van Regenmortel M, Fauquet C, Bishop D, Calisher C, Carsten E, Estes M et al (2002) Virus taxonomy. Seventh report of the International Committee for the Taxonomy of Viruses. Academic Press, New York
Bernard H, Burk R, Chen Z, van Doorslaer K, zur Hausen H, de Villiers E (2010) Classification of papillomaviruses (PVs) based on 189 PV types and proposal of taxonomic amendments. Virology 401:70–79
Yu JM, Ao YY, Zhao G, Li LL, Wang D, Duan ZJ (2013) Identification of a novel strain of human papillomavirus from children with diarrhea in china. Genome Announc 1(5)
Mokili JL, Dutilh BE, Lim YW, Schneider BS, Taylor T, Haynes MR et al (2013) Identification of a novel human papillomavirus by metagenomic analysis of samples from patients with febrile respiratory illness. PLoS ONE 8(3):e58404
Kocjan BJ, Hosnjak L, Seme K, Poljak M (2013) Complete genome sequence of a novel human betapapillomavirus, HPV-159. Genome Announc 1(3)
Phan TG, Vo NP, Aronen M, Jartti L, Jartti T, Delwart E (2013) Novel human gammapapillomavirus species in a nasal swab. Genome Announc 1(2):e0002213
Ramamoorthy S, Liu YT, Luo L, Miyai K, Lu Q, Carethers JM (2010) Detection of multiple human papillomavirus genotypes in anal carcinoma. Infect Agents Cancer 5:17
Bartholomew DA, Luff RD, Quigley NB, Curtis M, Olson MC (2011) Analytical performance of Cervista HPV 16/18 genotyping test for cervical cytology samples. J Clin Virol 51(1):38–43
Chaturvedi AK, Engels EA, Pfeiffer RM, Hernandez BY, Xiao W, Kim E et al (2011) Human papillomavirus and rising oropharyngeal cancer incidence in the United States. J Clin Oncol 29(32):4294–4301
Zhao G, Krishnamurthy S, Cai Z, Popov VL, TravassosdaRosa AP, Guzman H et al (2013) Identification of novel viruses using VirusHunter—an automated data analysis pipeline. PLoS ONE 8(10):78470
Titolo S, Pelletier A, Sauvé F, Brault K, Wardrop E, White PW et al (1999) Role of the ATP-binding domain of the human papillomavirus type 11 E1 helicase in E2-dependent binding to the origin. J Virol 73:5282–5293
Lehoux M, D’Abramo CM, Archambault J (2009) Molecular mechanisms of human papillomavirus-induced carcinogenesis. Public Health Genomics 12:268–280
Chouhy D, Bolatti EM, Piccirilli G, Sanchez A, Fernandez Bussy R, Giri AA (2012) Identification of human papillomavirus type 156, the prototype of a new human gammapapillomavirus species, by a generic and highly sensitive PCR strategy for long DNA fragments. J Gen Virol 94(Pt 3):524–533
de Villiers EM, Fauquet C, Broker TR, Bernard HU, zur Hausen H (2004) Classification of papillomaviruses. Virology 324(1):17–27
Acknowledgments
This work was supported by the National Natural Science Foundation of China (Grant No. 81290345) and by the China Mega-Project for Infectious Disease (Grant No. 2011ZX10004).
Conflict of interest
The authors declare that they have no competing interests.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yu, Jm., Zhao, G., Ao, Yy. et al. Complete genome sequence of a novel human papillomavirus identified by metagenomic analysis from a child with diarrhea in China. Arch Virol 160, 549–552 (2015). https://doi.org/10.1007/s00705-014-2252-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-014-2252-7