Abstract
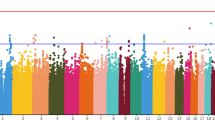
In recent years, several studies have investigated genetic polymorphisms of antipsychotic drug-metabolizing enzymes and receptors. However, most studies focused on drug response and very few have investigated the genetic influence on antipsychotic dosage. The aim of the present study is to test the association between antipsychotic dosages at genome-wide level. The current dosage of antipsychotic medications was collected from 79 schizophrenia patients. The dosage was standardized using three different methods: chlorpromazine equivalent (CPZe), defined daily dose (DDD), and percentage of maximum dose (PM %). The patients were then genotyped using the Illumina HumanOmni2.5-8 BeadChip Kit. All markers were screened for significance using linear regression, and the p values were visualized using a Manhattan plot. The genome-wide analysis showed that the top Single-Nucleotide Polymorphisms (SNPs) associated with dosage variation were rs981975 on chromosome 14 for CPZe, rs4470690 on chromosome 4 for PM %, and rs79323383 on chromosome 8 for DDD. However, no genome-wide significantly associated SNPs were identified. In this pilot sample, we found promising trends for pharmacodynamic targets associated with antipsychotic dosage. Therefore, studies combining large prescription databases may identify genetic predictors to adjust the dose of antipsychotic medication.










Similar content being viewed by others
References
Brandl EJ, Kennedy MD, James L, Müller MD, Daniel J (2014) Pharmacogenetics of antipsychotics. Canadian J Psychiatry 59(2):76–88
Caccia S (2011) Pharmacokinetics and metabolism update for some recent antipsychotics. Expert Opin Drug Metab Toxicol 7(7):829–846. doi:10.1517/17425255.2011.575061
Compendium of Pharmaceuticals and Specialties (2015) The Canadian Drug Reference for Health Professionals (CPS). In: Repchinsky C, Welbanks L, Hutsul J, Jovaisas B, Lewis G, Perrier H et al (eds). Canadian Pharmacists Association, Canada
Drago A, Giegling I, Schäfer M, Hartmann AM, Konte B, Friedl M, Serretti A, Rujescu D (2014) Genome-wide association study supports the role of the immunological system and of the neurodevelopmental processes in response to haloperidol treatment. Pharmacogenet Genomics 24(6):314–319
First MB, Spitzer RL, Gibbon M, Williams JBW (2002) Structured Clinical Interview for DSM-IV-TR Axis I Disorders, Research Version, Non-Patient Edition (SCID-I/NP). New York, New York State Psychiatric Institute.Biometrics Research
Gardner D, Murphy A, O’Donnell H, Centorrino F, Baldessarini R (2010) International consensus study of antipsychotic dosing. Am J Psychiatry 167:686–693
IBM Corp (2011) IBM SPSS Statistics for Windows, Version 20.0. Armonk, NY: IBM Corp
Kane J, Honigfeld G, Singer J, Meltzer H (1988) Clozapine for the Treatment-Resistant Schizophrenic: A Double-blind Comparison With Chlorpromazine. Arch Gen Psychiatry 45(9):789–796. doi:10.1001/archpsyc.1988.01800330013001
Kessler RC, Amminger GP, Aguilar-Gaxiola S, Alonso J, Lee S, Ustun TB (2007) Age of onset of mental disorders: a review of recent literature. Curr Opin Psychiatry. 20:359–364. doi:10.1097/YCO.0b013e32816ebc8c
McClay JL, Adkins DE, Aberg K, Bukszár J, Khachane AN, Keefe RS, Perkins DO, McEvoy JP, Stroup TS, Vann RE, Beardsley PM, Lieberman JA, Sullivan PF, van den Oord EJ (2011a) Genome-wide pharmacogenomic study of neurocognition as an indicator of antipsychotic treatment response in schizophrenia. Neuropsychopharmacology 36(3):616–626
McClay JL, Adkins DE, Aberg K, Stroup S, Perkins DO, Vladimirov VI, Lieberman JA, Sullivan PF, van den Oord EJ (2011b) Genome-wide pharmacogenomic analysis of response to treatment with antipsychotics. Mol Psychiatry 16(1):76–85
Pruim RJ, Welch RP, Sanna S, Teslovich TM, Chines PS et al (2010) LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics 26(18):2336–2337. doi:10.1093/bioinformatics/btq419
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D, Maller J, Sklar P, de Bakker PIW, Daly MJ, Sham PC (2007). PLINK: a toolset for whole-genome association and population-based linkage analysis. Am J Human Genet 81
R Core Team (2014) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. http://www.R-project.org/
van Os J, Kapur SS (2009) Lancet 374(9690):635–45. doi:10.1016/S0140-6736(09)60995-8
Van Sant SP, Buckley PF (2011) Pharmacotherapy for treatment-refractory schizophrenia. Expert Opin Pharmacother 12(3):411–434. doi:10.1517/14656566.2011.528200
World Health Organization (2013) WHO collaborating centre for drug statistics methodology. ATC/DDD index 2011. Available at http://www.whocc.no/atcddd
Wiersma D, Nienhuis FJ, Slooff CJ, Giel R (1998) Natural course of schizophrenic disorders: A 15-year followup of a dutch incidence cohort. Schizophr Bull 24(1):75–85. doi:10.1093/oxfordjournals.schbul.a033315
Acknowledgments
The authors like to thank the patients for investing their time in this study. Dr De Luca is supported by the CIHR Grant MOP-119332. Arthur T. Koga was under CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico—“National Counsel of Technological and Scientific Development”) scholarship during this research.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Koga, A.T., Strauss, J., Zai, C. et al. Genome-wide association analysis to predict optimal antipsychotic dosage in schizophrenia: a pilot study. J Neural Transm 123, 329–338 (2016). https://doi.org/10.1007/s00702-015-1472-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00702-015-1472-7