Abstract
Population-based studies have revealed 2–10% measles vaccine failure rate even after two vaccine doses. While the mechanisms behind this remain unknown, we hypothesized that host genetic factors are likely to be involved. We performed a genome-wide association study of measles specific neutralizing antibody and IFNγ ELISPOT response in a combined sample of 2872 subjects. We identified two distinct chromosome 1 regions (previously associated with MMR-related febrile seizures), associated with vaccine-induced measles neutralizing antibody titers. The 1q32 region contained 20 significant SNPs in/around the measles virus receptor-encoding CD46 gene, including the intronic rs2724384 (p value = 2.64 × 10−09) and rs2724374 (p value = 3.16 × 10−09) SNPs. The 1q31.1 region contained nine significant SNPs in/around IFI44L, including the intronic rs1333973 (p value = 1.41 × 10−10) and the missense rs273259 (His73Arg, p value = 2.87 × 10−10) SNPs. Analysis of differential exon usage with mRNA-Seq data and RT-PCR suggests the involvement of rs2724374 minor G allele in the CD46 STP region exon B skipping, resulting in shorter CD46 isoforms. Our study reveals common CD46 and IFI44L SNPs associated with measles-specific humoral immunity, and highlights the importance of alternative splicing/virus cellular receptor isoform usage as a mechanism explaining inter-individual variation in immune response after live measles vaccine.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Measles still remains a disease of public health concern in the developing world and well-developed countries with multiple outbreaks even among populations with high vaccine coverage. From 2010 to date, the European region registered 135,600 measles cases, and the US experienced 1381 measles cases in 27 states (Haralambieva et al. 2013, 2015; Poland and Jacobson 2012; Prevention 2015; Whitaker and Poland 2014). Several population-based studies have estimated that 2–10% of vaccine recipients do not develop or sustain measles-specific protective immunity after two doses of MMR vaccine (Bednarczyk et al. 2016; Haralambieva et al. 2011b, 2013; Poland and Jacobson 2012; Whitaker and Poland 2014). The mechanisms behind vaccine failure are unknown. This knowledge gap is an impediment to controlling future outbreaks or designing improved vaccine candidates.
Measles vaccine-induced humoral immunity is reported to have an extremely high heritability of 88.5% (Tan et al. 2001). We have performed a series of candidate genetic association studies delineating the effect of HLA alleles and single nucleotide polymorphisms on measles humoral and cellular immune responses, but thus far only approximately 30% of the inter-individual variation in immune response to this vaccine can be explained (Dhiman et al. 2007; Haralambieva et al. 2011a, c, 2013, 2015; Kennedy et al. 2012a; Ovsyannikova et al. 2011a, b, 2012).
We report the first GWAS study (on a sample of 2872 subjects) of measles vaccine-induced humoral and cellular immune response outcomes in children and younger adults, that identify significant SNP associations (in the CD46 and IFI44L genes) underlying the observed inter-individual variability in neutralizing antibody titers after vaccination.
Methods
Additional methods details are available in the Supplementary data.
Study subjects
We analyzed a large sample of 3191 healthy children, older adolescents, and healthy adults (age 11–40 years) consisting of three independent cohorts: a Rochester cohort (n = 1062); a San Diego cohort (n = 1071); and a US cohort (n = 1058). The demographic and clinical characteristics of these cohorts have been previously published (Haralambieva et al. 2011a, c; Kennedy et al. 2012a, b, c; Lambert et al. 2015; Ovsyannikova et al. 2011a, b, 2012).
The Institutional Review Boards of the Mayo Clinic (Rochester, MN) and the NHRC (San Diego, CA) approved the study, and written informed consent was obtained from each subject, i.e., from age-appropriate participants and the parents of all children who participated in the study.
Genotyping and immune outcomes
The genome-wide SNP typing was performed using the Infinium Omni 1M-Quad SNP array (Illumina; San Diego, CA) for the Rochester cohort, Illumina Human Omni2.5-8 BeadChip array for the US cohort, and Illumina Infinium HumanHap550v3_A or HumanHap650Yv3 BeadChip arrays for the San Diego cohort. Measles-specific neutralizing antibody and cytokine responses were quantified using a fluorescence-based plaque reduction microneutralization assay (PRMN) and ELISPOT/ELISA assays, as previously described (Haralambieva et al. 2011b). Our humoral immune response phenotype, the 50% end-point titer (Neutralizing Doze, ND50), was calculated using Karber’s formula and transformed into mIU/mL (using the 3rd WHO international measles antibody standard), as described previously (Haralambieva et al. 2011b). The variability of the PRMN assay, calculated as a coefficient of variation (CV) based on the log-transformed ND50 values of the third WHO standard, was 5.7%, as previously published (Haralambieva et al. 2011b).
Next generation sequencing (mRNA-Seq) and RT-PCR
Libraries were generated from total RNA (extracted from PBMCs of 30 subjects) using Illumina’s mRNA TruSeq (v1) kit and sequenced (paired end sequencing) on an Illumina HiSeq 2000 (Illumina; San Diego, CA) with Illumina’s TruSeq Cluster kit (v3-cBot-HS) and 51 Cycle Illumina TruSeq SBS Sequencing Kit (v3), as previously published (Haralambieva et al. 2016). One-Step RT-PCR system with Platinum Taq® DNA polymerase (Invitrogen, Carlsbad, CA) and primers allowing CD46 isoform/isoforms discrimination were used, as previously described (Wang et al. 2000).
Statistical methods
GWAS analysis
To achieve greatest power to detect SNPs associated with measles-specific immune response phenotypes, we pooled our data across genotyping platforms and the three cohorts. To perform the pooled analyses, we first accounted for the effects of potentially confounding factors that vary across ancestry/platform/cohort strata. After thoroughly evaluating the quality of the genotype data, we used the genetic data to define major ancestry groups. We then estimated eigenvectors within each ancestry group to account for the effects of population stratification within ancestry groups. Because the largest ancestry groups were Caucasian and African-American, we restricted our pooled analysis to these groups. After accounting for population stratification, we evaluated the covariates that were available within each of the ancestry-platform-cohort strata to determine if the covariates were associated with the phenotype, to regress out the effects of potential confounding factors. This produced residuals (adjusted traits) that were then used for the GWAS analyses. The immune response trait measles-specific IFNγ ELISPOT, as well as measles-specific secreted cytokines, were transformed by normal quantiles of the difference of the mean stimulated and mean unstimulated values. The immune response trait neutralizing antibody titer was transformed as the natural log of the PRMN mIU/mL value (for more details, refer to Supplemental Methods).
Analysis of mRNA-Seq data for differential exon usage
We tested for evidence of differential exon usage in the CD46 and IFI44L genes using the method of Anders et al., implemented in the DEXSeq package (version 1.16.10) in the R programing language (version 3.2.3) (Anders et al. 2012; Team 2009).
Molecular modeling
To evaluate the effect of differential splicing on the dynamics of the CD46 molecular structure, we generated homology models (Roy et al. 2010) of the extracellular domains (SCR1-4 and STP domains) for each isoform and analyzed each using Anisotropic Network Models (ANMs) (Atilgan et al. 2001; Chennubhotla and Bahar 2007; Yang et al. 2009) using multiple templates (see Supplemental Data for details). Interactions between CD46 and MV-H were modeled using the crystal structure PDB 3INB (Santiago et al. 2010) as a template wherein dimeric MV-H is bound symmetrically by the SCR1 and SCR2 domains of two CD46 molecules (see Supplemental Data for details).
Results
Genome-wide analysis results with humoral immunity
Genetic regions associated with variations in measles-specific antibody response after vaccination
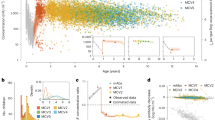
The demographic and immune characteristics of our study sample (n = 2872) are summarized in Table 1. We identified two independent gene regions on chromosome 1 associated with antibody response following measles vaccination (Fig. 1, Supplemental Fig. 1). As depicted on the locus zoom plot (Fig. 1a), the right region on chromosome 1 contained multiple SNPs (n = 20) in/around the MV receptor-encoding CD46 gene and region (1q32, bp 207917499-208025926, NCBI Build 37/hg19). Analyzing associations between antibody response and SNPs in the two other MV receptors on chromosome 1, SLAM (SLAMF1) and nectin-4 (NECTIN4/PVRL4), did not result in significant findings. The left region (Fig. 1c) on chromosome 1 (1p31.1, bp 79082772-79110518) contained nine significant SNPs in/around the interferon-induced, protein 44-like gene IFI44L.
Locus zoom plots of the chromosome 1 regions and effects of SNPs associated with measles-specific neutralizing antibody titers in the combined cohort (n = 2872, reduced to 2818 after excluding subjects with immune outcome data that failed QC). a, c Locus zoom plots of the 1q32(CD46, a) and 1q31.1 (IFI44L, c) regions associated with neutralizing antibody titer after measles vaccination (combined cohort, n = 2872, reduced to 2818 after excluding subjects with immune outcome data that failed QC). On the x axis SNPs are plotted by chromosomal location. The left y axis reflects the association (−log10 p value) with vaccine-induced measles-specific antibody titer, while the right y axis reflects recombination rates and LD (r 2 shades of gray [online version: color]) of each plotted SNP with the most significant SNP (designated by a black diamond). b, d Effect of top two CD46 SNPs (b) and IFI44L SNPs (d) on measles-specific antibody response (effect in the combined cohort is presented as Turkey box-and-whisker plots). On the x axis 0 designates subjects with homozygous major allele genotype, 1 designates heterozygous subjects and 2 designates subjects with homozygous minor allele genotype. On the y axis neutralizing antibody titer is presented as the natural log of the PRMN mIU/mL value. The top (bottom) of the box indicates the 75th (25th) percentiles, respectively, while the bold line within the box indicates the median, and the whiskers indicate 1.5 times the IQR (color figure online)
CD46 region SNPs associated with variations in measles-specific antibody response after vaccination
The genetic association signal from the 1q32 region was linked to two blocks (97 and 8 kb) of 20 genetic variants in and around the CD46 gene (seven intronic CD46 SNPs, four SNPs in the uncharacterized LOC101929385 [currently Gene ID 100128537, C1orf132 chromosome 1 open reading frame 132], and nine intergenic SNPs, including the previously reported rs1318653 (Feenstra et al. 2014), located between CD46 and CD34), which were in high linkage disequilibrium (LD) (Supplemental Fig. 2). The most significant CD46 SNPs, rs2724384 and rs2724374 (in high LD, r 2 = 97), lie in intron 1, and near the boundary of intron 8 (of the reference sequence ENST00000358170, RefSeq NM_002389), respectively. The minor alleles (G) of rs2724384 and rs2724374 were significantly associated (Table 2) with an allele dose-related decrease in measles-specific neutralizing antibody titers after vaccination (46–47% decrease in antibody titer in homozygous minor allele genotype subjects compared to homozygous major allele genotype subjects, Table 2; Fig. 1b). Due to the high LD (Supplemental Fig. 2), all significant CD46 SNPs displayed similar effects. Most of the 1q32 associations remained genome-wide significant (p < 5.0 × 10−08) in the subjects of Caucasian ancestry, where the most significant SNP was rs2724374 (p value = 4.88 × 10−09, Table 3).
IFI44L region SNPs associated with variations in measles-specific antibody response after vaccination
The 1p31.1 region signal was linked to a 21-kb block of 8 IFI44L SNPs and one intergenic SNP, in high LD (Supplemental Fig. 2). The two most significant IFI44L SNPs, rs1333973 and rs273259 (Table 2) were located in IFI44L intron 2 (boundary) and IFI44L exon 2, respectively. The missense SNP rs273259 (His73Arg, Ensembl transcript ENST00000370751), as well as the other significant SNPs, demonstrated an allele dose-related decrease in measles-specific neutralizing antibody titers (Table 2; Fig. 1d). Three of the nine SNPs remained genome-wide significant (p < 5.0 × 10−08) in the subjects of Caucasian ancestry (Table 3).
To determine if there were multiple SNPs in each of the CD46 and IFI44L regions associated with the neutralizing antibody, with the effects of the SNPs adjusted for each other, we used elastic net to select SNPs. An advantage of elastic net is that it can select highly correlated SNPs, although it cannot give p values for selected SNPs. Hence, after selection, we evaluated the selected SNPs in linear regression models. This resulted in the selection of two SNPs in the CD46 region and three SNPs in the IFI44L region, based on the Caucasian subjects. For the CD46 region, we found rs2724374 to be the most significant and the primary driving SNP (p = 4.3 × 10−8), with rs11806810 having much less association (p = 0.02) when the effects of the SNPs were adjusted for each other. For the IFI44L region, the SNPs rs12026737, rs1333973, and rs273259 were selected. However, rs273259 and rs1333973 were highly correlated (r 2 = 0.99), making it difficult to disentangle their effects and fit both in a regression model. When choosing rs1333973 (top significant SNP) over rs273259, we found rs1333973 (p = 2 × 10−8) and rs12026737 (p = 6.0 × 10−4) to have strong statistical associations, with effects adjusted for each other. Furthermore, the CD46 and IFI44L regions have associations that are independent of each other (since when the aforementioned SNPs were modeled together, the results were not significantly different from what was obtained when the regions were modeled separately).
Genome-wide analysis results with measles vaccine cellular immunity
The GWAS analyses did not reveal significant SNP associations with cellular immunity after vaccination, as measured by MV-specific IFNγ ELISPOT (Supplemental Table 1, Supplemental Fig. 3). Analyses of all chromosome 1 SNPs with MV-specific secreted cytokines in 625 Caucasian subjects (for whom we had available cytokine data) demonstrate suggestive associations between CD46 SNPs (including rs2724374 and rs2724384) and the secretion of IFNα; however, an allele-dose-dependency of IFNα secretion was not noted (Supplemental Table 2).
Analysis of differential usage of exons using mRNA-Seq data
In the CD46 analyses, we observed a highly significant per-exon estimate (q = 2.96E−07) with lower exon B (in the CD46 STP region) expression (exon skipping) in subjects with rs2724374 minor G allele (Fig. 2a). These results were further confirmed by RT-PCR analysis of common CD46 isoforms in PBMCs. In this analysis, the predominant “lower band” CD46 isoform phenotype (C2, associated with exon B skipping) was clearly more pronounced in the homozygous minor G allele genotype subjects (Fig. 2b). We also observed differential exon usage based on IFI44L rs1333973/rs273259 genotypes (Fig. 2c).
Differential exon/isoform CD46 and IFI44L usage. a Estimated exon usage in the CD46 gene comparing individuals with at least one minor allele (G for rs2724374, i.e., all the heterozygous subjects plus the one homozygous minor allele subject combined, total n = 9) (light gray lines [online version: blue]) to individuals that are homozygous major (T for rs2724374) allele, n = 19 (dark gray lines [online version: red]). Even with a relatively small number of subjects possessing the minor allele genotype, differential exon usage with a highly significant p value was observed for the STP exon B (genomic ID 207941124-207941168) (p = 2.96E−07). b RT-PCR analysis of common CD46 isoforms was performed in PBMCs of rs2724374 homozygous major allele genotype individuals (n = 10) compared to homozygous minor allele genotype individuals (n = 10) and heterozygous individuals (n = 10). The presented figure is representative of the patterns observed in all 30 subjects (10 subjects per genotype group) with the experiment replicated twice. c Estimated exon usage in IFI44L rs1333973/rs273259 homozygous minor allele genotype subjects (A for rs1333973, n = 5 [light gray/blue]) vs. homozygous major allele subjects (T for rs1333973, n = 11 [dark gray/red]). The results demonstrate significant per-exon estimates for several IFI44L exons, the most significant being exon 2 (genomic ID 79093591-79094078) (q = 2.37E−150) (color figure online)
Simulation of the structural differences between CD46 isoforms (with or without the STP exon B)
We have generated structural models of the predominant CD46 isoforms (extracellular portion): isoforms with the STP exons B and C (i.e., BC1 and BC2) and those skipping exon B (i.e., C1 and C2) (see Figs. 3, 4, Supplemental Fig. 4). The longest isoform, ABC1, generates a high-quality homology model (Supplemental Fig. 4). Interestingly, the protein sequences encoded by exons A (exon 7) and B (exon 8) have their N- and C-terminal ends close to each other. The amino acids encoded by exon A (exon 7) make a short beta strand within the STP domain. Deletion of these residues by exon skipping removes this strand from the beta-sheet, but the architecture of the domain is not significantly altered. Exon B (8) encodes amino acids making up two additional strands. In the isoforms C1 and C2, only two strands encoded by exons 9 and 10 remain. Computing the dynamics of each isoform’s atomic model, smaller STP domain isoforms exhibited greater flexibility and less collective motion (i.e., in the C1/C2 isoforms when compared to BC1/BC2 of the common isoforms). Monitoring the domain–domain angles about the SCR4-STP hinge as each structure is deformed about its low-frequency normal modes, the shorter isoform/isoforms exhibit a greater range of flexibility (Supplemental Fig. 4). As illustrated in Fig. 4, based on data from our molecular models, we propose that the differential isoform flexibility may influence the rate of formation of the CD46-MV-hemagglutinin (H) encounter complex, and/or influence the propensity for MV-H to undergo the conformational change necessary to trigger MV fusion protein.
Structure of CD46. The extracellular portion of CD46 consists of four N-glycosylated conserved short consensus repeats SCR1-4 (SCR1 and SCR2 containing binding sites for MV); a STP region that is O-glycosylated (encoded by exons 7, 8 and 9, designated as A, B and C); and a region of unknown function (U), followed by a hydrophobic transmembrane segment (H), basic amino acid anchor (A), and a cytoplasmic tail (CYT1 of 16 amino acids if exon 13 is present, or CYT2 of 23 amino acids if exon 13 is alternatively spliced). Of the 14 known CD46 isoforms resulting from alternative splicing, four are commonly found in most human tissues and are designated based on the present STP exon/exons and the cytoplasmic tail: BC1 and BC2 (with B and C exons/domains in the STP and with either CYT1 or CYT2), and C1 and C2 (with C exon/domain in the STP and with either CYT1 or CYT2). The effect of CD46 rs2724374 on CD46 isoform prevalence (exon B presence or skipping), interaction between CD46 and MV, and immune response following measles vaccination is also summarized
Flexibility of CD46 isoforms and impact on MV-CD46 interactions and MV fusion. a CD46 is a cell surface receptor affected by isoform differences. Due in part to its elongated structure, the extracellular domains will naturally exhibit flexibility within the physiologic environment (represented as rotational blurring of the reference isoform ABC1/ABC2). SCR domain residues are colored in light gray (online version: yellow). The modeled segment of CD46 includes the extracellular domains and is shown in cartoon representation. b Molecular modeling of the extracellular portion of BC1 (BC2) and C1 (C2) isoforms (the four most common isoforms) through SCR4, followed by generation of mechanics-based models (ANM; see “Methods”), indicates a difference in the intrinsic flexibility between the isoforms (see Supplemental Fig. 4 for details). The amino acids encoded by exons 8–10 are indicated by shades of gray (online version: different colors). We show representative motion from our ANM models for the BC1 and C1 isoforms by showing multiple conformational states superimposed. c CD46 on the cell surface interacts with MV-H on the virion surface. Two CD46 molecules interacting with a MV-H dimer are depicted, as described previously (Santiago et al. 2010). Using ANM, we quantified the intrinsic flexibility of the MV-H dimer. The dominant motion is an anti-correlated twisting or ratcheting of the monomers with respect to one another. Projection the effect of this motion onto the bound CD46 molecules is represented by black arrows on each residue. As the C-terminus of CD46 is anchored in the cell membrane, activation of this motion would require flexibility of the CD46 molecule—flexibility that may differ by isoform (panel b). Presented is a second view of the motion rotated 90°, omitting the cartoon representation for clarity—only the motion-vectors are shown to emphasize the twisting of the MV-H dimer and its effect on CD46. d Proposed molecular mechanism: MV-H and the fusion protein MV-F are normally associated with one another on the virus surface. Proteins are represented by smoothed molecular surfaces; see Supplemental Methods. The encounter complex between CD46 and MV-H leads to MV-F disassociation. The disassociation is influenced by the MV-H conformational change and motion in the context of the CD46-H dimer complex (Navaratnarajah et al. 2011). The free MV-F undergoes a substantial conformational change, the molecular details of which are not fully resolved, leading to bridging between the virus and target cell. We believe that the degree of flexibility exhibited by CD46 may influence (1) the ease with which the complex may undergo conformational changes (motion in the context of MV-H-CD46 complex) leading to MV-F triggering and fusion, and (2) the rate of encounter complex formation with MV-H (color figure online)
Discussion
This work is the first GWAS to reveal significant associations of two distinct chromosome 1 regions (CD46 and IFI44L) with variations in neutralizing antibody response after measles vaccination.
The important implications of our findings are supported by the genome-wide identification of genetic variants in the same two loci (in an independent state-of-the-art GWAS study) as risk variants for the development of adverse events/febrile seizures following MMR vaccination, but not for MMR-unrelated febrile seizures (suggesting the influence of these loci on measles vaccine virus entry/propagation) (Feenstra et al. 2014).
Most of the significant SNP-association signals with measles-specific antibody response were from a region in and around the CD46 gene on 1q32. The encoded glycoprotein CD46 (structure is summarized in Fig. 3) is ubiquitously expressed and serves as a regulator of complement activation (protecting the cells from complement and antibody-mediated lysis). CD46 also serves as a cellular receptor for measles virus attenuated (vaccine) strains, as well as for other pathogens (group B and D adenoviruses, human herpesvirus 6, bovine viral diarrhea virus, pathogenic Neisseria and Streptococcus pyogenes). (Cattaneo 2004) Four CD46 isoforms, resulting from alternative splicing, are commonly found in most human tissues and are designated based on the present STP exon/exons and the cytoplasmic tail: BC1 and BC2 (with B and C exons/domains in the STP and with either CYT1 or CYT2), and C1 and C2 (with C exon/domain in the STP and with either CYT1 or CYT2) (Liszewski et al. 1994; Post et al. 1991; Russell et al. 1992) (Fig. 3).
Our significant GWAS findings in the CD46 region consisted only of non-coding SNPs in high LD; the most significant were rs2724384 in CD46 intron 1, and rs2724374 in CD46 intron 8. The previously observed (Feenstra et al. 2014) genetic association of intergenic rs1318653 with MMR-related febrile seizures is in the same region and was also significant in our genome-wide association study with measles-specific neutralizing antibody titers (p value = 2.94 × 10−08, Table 2), although this association exhibited a slightly weaker signal in the subset analysis of subjects of Caucasian ancestry (p value = 1.04 × 10−07, Table 3). Feenstra et al. also reported the major CD46 rs2724384 allele A as a risk allele for MMR-related febrile seizures (Feenstra et al. 2014), which relates to higher measles vaccine-induced antibody titers in our GWAS and in three other candidate gene studies. (Clifford et al. 2012; Dhiman et al. 2007; Haralambieva et al. 2015; Ovsyannikova et al. 2011a) Furthermore, CD46 rs2724384 has been associated with overall CD46 gene expression and isoform abundance in lymphoblastoid cell lines (Lappalainen et al. 2013), and with variations in measles-specific IL-6, TNFα and IFNα secretion after in vitro viral stimulation of human PBMCs (Ovsyannikova et al. 2011a). In this study, both of our top CD46 GWAS hits (rs2724384 and rs2724374) exhibited suggestive associations with IFNα (see Supplemental Table 2).
The second CD46 genetic variant with plausible functional consequences is rs2724374 (located in intron 8, near the intron–exon boundary with exon 8/STP B), which is the most significant SNP in our analysis of subjects of Caucasian ancestry and is identified by elastic net modeling as being predictive of neutralizing antibody response. CD46 rs2724374 has been reported as a genetic variant, which is highly correlated with the splicing/skipping of exon B (i.e., a sQTL/splicing quantitative trait locus) in a study assessing the splicing patterns of 250 exons and their associations with DNA polymorphisms (Hull et al. 2007). Similarly, a study assessing sQTLs from RNA-Seq data reported CD46 rs2724374 as one of the most significant sQTLs (p = 3.55 × 10−11) associated with alternative splicing (i.e., the skipping of CD46 exon B) (Zhao et al. 2013), a finding validated by our differential exon usage analysis of mRNA-Seq data and RT-PCR analysis of CD46 isoforms in PBMCs. Thus, the minor allele G of CD46 rs2724374 is significantly associated (in a dose–response dependent manner) with lower measles-specific neutralizing antibody titer after vaccination (approximately 45% reduction in antibody titer), and with the skipping of exon B to preferentially yielding CD46 isoforms with a shorter (and less O-glycosylated) STP region (i.e., preferentially yielding C1/C2 vs. BC1/BC2 isoforms). Although some tissues preferentially express specific CD46 isoforms (sperm, brain, salivary gland, kidney, placenta), the abundance ratio of CD46 isoforms in most human tissues (e.g., PBMCs) is considered to be constant for each individual (Liszewski et al. 1994; Post et al. 1991; Russell et al. 1992). These CD46 genetic and phenotypic inter-individual variations and their implications for host-pathogen interactions and vaccine/pathogen-induced immunity still remain unknown.
Since innate and adaptive immune responses rely on recognition, virus entry and propagation into susceptible cells, it is tempting to speculate about the mechanisms by which CD46 isoform usage can affect MV binding/fusion and propagation at the site of injection and the associated lymphoid tissue during measles vaccination and immune response priming. It is generally accepted that all CD46 isoforms can serve as MV receptors (primarily for attenuated MV strains) and confer susceptibility to infection (Manchester et al. 1994); however, several studies report differences in MV binding and fusion in CD46 BC1/BC2 vs. C1/C2 isoforms, where longer BC isoforms support superior virus binding and shorter C isoforms support superior fusion competence (Buchholz et al. 1996a, b; Iwata et al. 1994). It is likely that the complex dynamics of interactions between MV H dimers/tetramers, F trimers and cross-linked CD46 molecules at the virus-cell surface interface (Navaratnarajah et al. 2011; Persson et al. 2010) are dependent in part on the varying flexibility conferred by different CD46 isoforms. Our molecular modeling (Fig. 4 and Supplemental Fig. 4) provides evidence for differences in the flexibilities of common CD46 isoforms, including the STP exon B (BC1/BC2 exhibiting decreased flexibility) and those excluding exon B (C1/C2 exhibiting increased flexibility). We hypothesize that the increased flexibility for the C1/C2 isoforms relative to BC1/BC2 isoforms allows increased motion between the components of the CD46-H dimer complex [needed for F triggering, as proposed in the model by Navaratnarajah et al. (2011) and Persson et al. (2010)] and increased fusogenicity. It is also possible that the increased flexibility of the C1/C2 isoforms (relative to BC1/BC2) may increase the encounter rate between CD46 and H protein. Thus, our proposed mechanistic hypothesis (informed by molecular modeling) is in concert with—and adds to—the prior literature, and suggests potential molecular mechanisms linking the top GWAS CD46 hits to differences in splicing/CD46 isoform flexibility with potential effects on MV response.
The CYT1 and CYT2-containing CD46 isoforms (generated by alternative splicing of exon 13) are reported to differentially co-stimulate T helper 1 effector cells, affect T helper 1 switching to IL-10 producing T regulatory cells, and modulate cellular immunity, inflammation, B cell responses and signal transduction (Fuchs et al. 2009; Marie et al. 2002; Wang et al. 2000). Although the differential usage of CYT1 vs. CYT2-containing isoforms cannot be inferred from our data, the involvement of such mechanisms in regulating vaccine-induced immunity cannot be excluded. Interestingly, we did not observe significant associations between polymorphisms in the other known MV receptor genes (SLAM and nectin-4) and vaccine-induced humoral immunity. This finding suggests the preferential usage of CD46 (over other receptors) by MV vaccine strains and/or insufficient expression of SLAM and Nectin-4 receptors at the site of injection and immune response priming during vaccination.
Even more interesting is the association of the 1p31.1 IFI44L genetic locus with measles-specific antibody titers, since the function of the encoded protein is largely unknown. Among the top significant SNPs in this region (grouped in a 21-kb LD block) are the intronic rs1333973 and the coding missense rs273259 (Tables 2, 3); the latter has been reported to be a genetic variant significantly associated with febrile seizures after MMR vaccination (Feenstra et al. 2014). The rs273259 risk allele (A) for MMR-related febrile seizures was associated with increased measles-specific antibody titer in our study (while the minor allele G was associated with decreased antibody titer). In addition, rs273259 allele A was previously found to correlate with both the reduced expression of IFI44L exon 2 and differences in IFI44L isoform/isoforms abundance (Lappalainen et al. 2013), a finding also confirmed by our mRNA-Seq differential exon usage analysis (Fig. 2c). In their study, Feenstra et al. did not observe direct and/or differential antiviral activity of IFI44L rs273259 allelic variant proteins against MV in human fibroblasts lacking STAT1 (Feenstra et al. 2014). It is likely that IFI44L requires a specific microenvironment (e.g., partners, signaling events, cell/tissue-specific milieu, etc.) to exert its antiviral action or its function is associated with specific isoforms, as suggested by our data and other studies [i.e., SNP rs1333973 has been reported as a significant sQTL for IFI44L (Coulombe-Huntington et al. 2009; Fraser and Xie 2009; Zhao et al. 2013)].
The proteins encoded by IFI44L and its partner IFI44 are both stimulated by interferon type I and thus are likely to be involved in innate immunity. A screen of more than 380 interferon stimulated genes/ISGs for in vitro antiviral activity against different viruses identified IFI44L as an important antiviral effector of type I interferon response against hepatitis C virus (Schoggins et al. 2011). Importantly, a study assessing susceptibility to viral myocarditis in mice (by Coxsackievirus B3) noted that genetic variants in the H28 (IFI44L) locus are likely involved in infection susceptibility (Wiltshire et al. 2011). These data from the literature suggest that IFI44L is an important effector of innate antiviral immunity with a plausible role in immune response priming and protection against disease.
The strengths of our study include the use of a relatively large combined sample of three well-characterized cohorts (from different geographic areas) and the detailed and reliable demographic, clinical, immune phenotyping and genomic data to classify study participants into genetically defined ancestry categories. Despite the popularity of the discovery/replication study design, we analyzed available cohorts together [as this is the currently accepted approach (McCarthy et al. 2008; Skol et al. 2006), and taking advantage of the full set of genotyping data], which resulted in greater power and provided more accurate estimates of the magnitude of SNP associations and genomic localization. Although our significant findings (i.e., the involvement of CD46 and IFI44L in the host response to measles vaccine) are supported by a different state-of-the-art GWAS study [Feenstra et al. GWAS study of febrile seizures (Feenstra et al. 2014)], functional mechanistic studies are warranted to elucidate the underlying molecular mechanisms and the link between CD46 and IFI44L genetic variants and humoral immunity to MV.
In conclusion, our study identified common genetic variants associated with inter-individual variations in measles-specific antibody response following MMR vaccination. Ultimately, our study significantly advances the science of measles vaccine immunology/immunogenetics and provides knowledge of the most critical genomic features influencing measles vaccine immune response and their potential molecular mechanisms—a foundational information for the development of better measles vaccine candidates and vaccination approaches. Our study also provides more general insights and a proof of concept for the critical role of genomic variability in virus/pathogen cellular receptors and receptor isoform usage for the development of humoral immunity following vaccination.
References
Anders S, Reyes A, Huber W (2012) Detecting differential usage of exons from RNA-seq data. Genome Res 22:2008–2017. doi:10.1101/gr.133744.111
Atilgan AR, Durell SR, Jernigan RL, Demirel MC, Keskin O, Bahar I (2001) Anisotropy of fluctuation dynamics of proteins with an elastic network model. Biophys J 80:505–515. doi:10.1016/S0006-3495(01)76033-X
Bednarczyk RA, Orenstein WA, Omer SB (2016) Estimating the number of measles-susceptible children and adolescents in the United States Using data from the national immunization survey-teen (NIS-teen). Am J Epidemiol 184:148–156. doi:10.1093/aje/kwv320
Buchholz CJ, Gerlier D, Hu A, Cathomen T, Liszewski MK, Atkinson JP, Cattaneo R (1996a) Selective expression of a subset of measles virus receptor-competent CD46 isoforms in human brain. Virology 217:349–355
Buchholz CJ, Schneider U, Devaux P, Gerlier D, Cattaneo R (1996b) Cell entry by measles virus: long hybrid receptors uncouple binding from membrane fusion. Virology 70:3716–3723
Cattaneo R (2004) Four viruses, two bacteria, and one receptor: membrane cofactor protein (CD46) as pathogens’ magnet. J Virol 78:4385–4388
Chennubhotla C, Bahar I (2007) Signal propagation in proteins and relation to equilibrium fluctuations. PLoS Comput Biol 3:1716–1726. doi:10.1371/journal.pcbi.0030172
Clifford HD, Hayden CM, Khoo SK, Zhang G, Le Souef PN, Richmond P (2012) CD46 measles virus receptor polymorphisms influence receptor protein expression and primary measles vaccine responses in naive Australian children. Clin Vaccine Immunol 19:704–710
Coulombe-Huntington J, Lam KC, Dias C, Majewski J (2009) Fine-scale variation and genetic determinants of alternative splicing across individuals. PLoS Genet 5:e1000766. doi:10.1371/journal.pgen.1000766
Dhiman N, Cunningham JM, Jacobson RM, Vierkant RA, Wu Y, Ovsyannikova IG, Pankratz VS, Poland GA (2007) Variations in measles vaccine-specific humoral immunity by polymorphisms in SLAM and CD46 measles virus receptors. J Allergy Clin Immunol 120:666–672
Feenstra B, Pasternak B, Geller F, Carstensen L, Wang T, Huang F, Eitson JL, Hollegaard MV, Svanstrom H, Vestergaard M, Hougaard DM, Schoggins JW, Jan LY, Melbye M, Hviid A (2014) Common variants associated with general and MMR vaccine-related febrile seizures. Nat Genet 46:1274–1282. doi:10.1038/ng.3129
Fraser HB, Xie X (2009) Common polymorphic transcript variation in human disease. Genome Res 19:567–575. doi:10.1101/gr.083477.108
Fuchs A, Atkinson JP, Fremeaux-Bacchi V, Kemper C (2009) CD46-induced human Treg enhance B-cell responses. Eur J Immunol 39:3097–3109. doi:10.1002/eji.200939392
Haralambieva IH, Ovsyannikova IG, Kennedy RB, Vierkant RA, Pankratz SV, Jacobson RM, Poland GA (2011a) Associations between single nucleotide polymorphisms and haplotypes in cytokine and cytokine receptor genes and immunity to measles vaccination. Vaccine 29:7883–7895
Haralambieva IH, Ovsyannikova IG, O’Byrne M, Pankratz VS, Jacobson RM, Poland GA (2011b) A large observational study to concurrently assess persistence of measles specific B-cell and T-cell immunity in individuals following two doses of MMR vaccine. Vaccine 29:4485–4491
Haralambieva IH, Ovsyannikova IG, Umlauf BJ, Vierkant RA, Pankratz SV, Jacobson RM, Poland GA (2011c) Genetic polymorphisms in host antiviral genes: associations with humoral and cellular immunity to measles vaccine. Vaccine 29:8988–8997
Haralambieva IH, Ovsyannikova IG, Pankratz VS, Kennedy RB, Jacobson RM, Poland GA (2013) The genetic basis for interindividual immune response variation to measles vaccine: new understanding and new vaccine approaches. Expert Rev Vaccines 12:57–70. doi:10.1586/erv.12.134
Haralambieva IH, Kennedy RB, Ovsyannikova IG, Whitaker JA, Poland GA (2015) Variability in humoral immunity to measles vaccine: new developments. Trends Mol Med 21:789–801. doi:10.1016/j.molmed.2015.10.005
Haralambieva IH, Zimmermann MT, Ovsyannikova IG, Grill DE, Oberg AL, Kennedy RB, Poland GA (2016) Whole transcriptome profiling identifies CD93 and other plasma cell survival factor genes associated with measles-specific antibody response after vaccination. PLoS One 11:e0160970. doi:10.1371/journal.pone.0160970
Hull J, Campino S, Rowlands K, Chan MS, Copley RR, Taylor MS, Rockett K, Elvidge G, Keating B, Knight J, Kwiatkowski D (2007) Identification of common genetic variation that modulates alternative splicing. PLoS Genet 3:e99. doi:10.1371/journal.pgen.0030099
Iwata K, Seya T, Ueda S, Ariga H, Nagasawa S (1994) Modulation of complement regulatory function and measles virus receptor function by the serine-threonine-rich domains of membrane cofactor protein (CD46). Biochem J 304(Pt 1):169–175
Kennedy RB, Ovsyannikova IG, Haralambieva IH, O’Byrne MM, Jacobson RM, Pankratz VS, Poland GA (2012a) Multigenic control of measles vaccine immunity mediated by polymorphisms in measles receptor, innate pathway, and cytokine genes. Vaccine 30:2159–2167
Kennedy RB, Ovsyannikova IG, Pankratz VS, Haralambieva IH, Vierkant RA, Jacobson RM, Poland GA (2012b) Genome-wide genetic associations with IFNgamma response to smallpox vaccine. Hum Genet 131:1433–1451
Kennedy RB, Ovsyannikova IG, Shane PV, Haralambieva IH, Vierkant RA, Poland GA (2012c) Genome-wide analysis of polymorphisms associated with cytokine responses in smallpox vaccine recipients. Hum Genet 131:1403–1421
Lambert ND, Haralambieva IH, Kennedy RB, Ovsyannikova IG, Pankrantz VS, Poland GA (2015) Polymorphisms in HLA-DPB1 are associated with differences in rubella-specific humoral immunity after vaccination. J Infect Dis 211:898–905
Lappalainen T, Sammeth M, Friedlander MR, t Hoen PA, Monlong J, Rivas MA, Gonzalez-Porta M, Kurbatova N, Griebel T, Ferreira PG, Barann M, Wieland T, Greger L, van Iterson M, Almlof J, Ribeca P, Pulyakhina I, Esser D, Giger T, Tikhonov A, Sultan M, Bertier G, MacArthur DG, Lek M, Lizano E, Buermans HP, Padioleau I, Schwarzmayr T, Karlberg O, Ongen H, Kilpinen H, Beltran S, Gut M, Kahlem K, Amstislavskiy V, Stegle O, Pirinen M, Montgomery SB, Donnelly P, McCarthy MI, Flicek P, Strom TM, Lehrach H, Schreiber S, Sudbrak R, Carracedo A, Antonarakis SE, Hasler R, Syvanen AC, van Ommen GJ, Brazma A, Meitinger T, Rosenstiel P, Guigo R, Gut IG, Estivill X, Dermitzakis ET (2013) Transcriptome and genome sequencing uncovers functional variation in humans. Nature 501:506–511. doi:10.1038/nature12531
Liszewski MK, Tedja I, Atkinson JP (1994) Membrane cofactor protein (CD46) of complement. Processing differences related to alternatively spliced cytoplasmic domains. J Biol Chem 269:10776–10779
Manchester M, Liszewski MK, Atkinson JP, Oldstone MBA (1994) Multiple isoforms of CD46 (membrane cofactor protein) serve as receptors for measles virus. Proc Natl Acad Sci USA 91:2161–2165
Marie JC, Astier AL, Rivailler P, Rabourdin-Combe C, Wild TF, Horvat B (2002) Linking innate and acquired immunity: divergent role of CD46 cytoplasmic domains in T cell induced inflammation. Nat Immunol 3:659–666. doi:10.1038/ni810
McCarthy MI, Abecasis GR, Cardon LR, Goldstein DB, Little J, Ioannidis JP, Hirschhorn JN (2008) Genome-wide association studies for complex traits: consensus, uncertainty and challenges. Nat Rev Genet 9:356–369
Navaratnarajah CK, Oezguen N, Rupp L, Kay L, Leonard VH, Braun W, Cattaneo R (2011) The heads of the measles virus attachment protein move to transmit the fusion-triggering signal. Nat Struct Mol Biol 18:128–134
Ovsyannikova IG, Haralambieva IH, Vierkant RA, O’Byrne MM, Jacobson RM, Poland GA (2011a) The association of CD46, SLAM, and CD209 cellular receptor gene SNPs with variations in measles vaccine-induced immune responses—a replication study and examination of novel polymorphisms. Hum Hered 72:206–223
Ovsyannikova IG, Haralambieva IH, Vierkant RA, Pankratz VS, Poland GA (2011b) The role of polymorphisms in toll-like receptors and their associated intracellular signaling genes in measles vaccine immunity. Hum Genet 130:547–561
Ovsyannikova IG, Pankratz VS, Vierkant RA, Jacobson RM, Poland GA (2012) Consistency of HLA associations between two independent measles vaccine cohorts: a replication study. Vaccine 30:2146–2152
Persson BD, Schmitz NB, Santiago C, Zocher G, Larvie M, Scheu U, Casasnovas JM, Stehle T (2010) Structure of the extracellular portion of CD46 provides insights into its interactions with complement proteins and pathogens. PLoS Pathog 6:e1001122. doi:10.1371/journal.ppat.1001122
Poland GA, Jacobson RM (2012) The re-emergence of measles in developed countries: time to develop the next-generation measles vaccines? Vaccine 30:103–104
Post TW, Liszewski MK, Adams EM, Tedja I, Miller EA, Atkinson JP (1991) Membrane cofactor protein of the complement system: alternative splicing of serine/threonine/proline-rich exons and cytoplasmic tails produces multiple isoforms that correlate with protein phenotype. J Exp Med 174:93–102
Prevention CfDCa (2015) Measles Cases and Outbreaks. http://www.cdc.gov/measles/cases-outbreaks.html. http://www.cdc.gov/measles/cases-outbreaks.html. Accessed 12 Feb 2015
Roy A, Kucukural A, Zhang Y (2010) I-TASSER: a unified platform for automated protein structure and function prediction. Nat Protoc 5:725–738. doi:10.1038/nprot.2010.5
Russell SM, Sparrow RL, McKenzie IFC, Purcell DFJ (1992) Tissue-specific and allelic expression of the complement regulator CD46 is controlled by alternative splicing. Eur J Immunol 22:1513–1518
Santiago C, Celma ML, Stehle T, Casasnovas JM (2010) Structure of the measles virus hemagglutinin bound to the CD46 receptor. Nat Struct Mol Biol 17:124–129. doi:10.1038/nsmb.1726
Schoggins JW, Wilson SJ, Panis M, Murphy MY, Jones CT, Bieniasz P, Rice CM (2011) A diverse range of gene products are effectors of the type I interferon antiviral response. Nature 472:481–485
Skol AD, Scott LJ, Abecasis GR, Boehnke M (2006) Joint analysis is more efficient than replication-based analysis for two-stage genome-wide association studies. Nat Genet 38:209–213
Tan PL, Jacobson RM, Poland GA, Jacobsen SJ, Pankratz SV (2001) Twin studies of immunogenicity—determining the genetic contribution to vaccine failure. Vaccine 19:2434–2439
Team RDC (2009) R: a language for statistical computing. R Foundation for Statistical Computing, Vienna
Wang G, Liszewski MK, Chan AC, Atkinson JP (2000) Membrane cofactor protein (MCP; CD46): isoform-specific tyrosine phosphorylation. J Immunol 164:1839–1846
Whitaker JA, Poland GA (2014) Measles and mumps outbreaks in the United States: think globally, vaccinate locally. Vaccine 32:4703–4704. doi:10.1016/j.vaccine.2014.06.088
Wiltshire SA, Leiva-Torres GA, Vidal SM (2011) Quantitative trait locus analysis, pathway analysis, and consomic mapping show genetic variants of Tnni3k, Fpgt, or H28 control susceptibility to viral myocarditis. J Immunol 186:6398–6405. doi:10.4049/jimmunol.1100159
Yang L, Song G, Jernigan RL (2009) Protein elastic network models and the ranges of cooperativity. Proc Natl Acad Sci USA 106:12347–12352. doi:10.1073/pnas.0902159106
Zhao K, Lu ZX, Park JW, Zhou Q, Xing Y (2013) GLiMMPS: robust statistical model for regulatory variation of alternative splicing using RNA-seq data. Genome Biol 14:R74. doi:10.1186/gb-2013-14-7-r74
Acknowledgements
We thank the Mayo Clinic Vaccine Research Group staff and the study participants. We wish to recognize Julie M. Cunningham and the Mayo Advanced Genomic Technology Center for the genotyping and next generation sequencing efforts, and Nathaniel D. Warner (Division of Biomedical Statistics and Informatics, Mayo Clinic Department of Health Science Research) for his programming assistance and contribution to statistical analysis. We thank Caroline L. Vitse for her editorial assistance with this manuscript. Research reported in this publication was supported by the National Institute of Allergy And Infectious Diseases of the National Institutes of Health under Award Number R01AI033144 and R37AI048793. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Dr. Poland is the chair of a Safety Evaluation Committee for novel investigational vaccine trials being conducted by Merck Research Laboratories. Dr. Poland offers consultative advice on vaccine development to Merck & Co. Inc., CSL Biotherapies, Avianax, Dynavax, Novartis Vaccines and Therapeutics, Emergent Biosolutions, Adjuvance, Microdermis, Seqirus, NewLink, Protein Sciences, GSK Vaccines, and Sanofi Pasteur. Drs. Poland and Ovsyannikova hold three patents related to measles and vaccinia peptide research. Dr. Kennedy has received funding from Merck Research Laboratories to study waning immunity to measles and mumps after immunization with MMR-II®. These activities have been reviewed by the Mayo Clinic Conflict of Interest Review Board and are conducted in compliance with Mayo Clinic Conflict of Interest policies. This research has been reviewed by the Mayo Clinic Conflict of Interest Review Board and was conducted in compliance with Mayo Clinic Conflict of Interest policies.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Haralambieva, I.H., Ovsyannikova, I.G., Kennedy, R.B. et al. Genome-wide associations of CD46 and IFI44L genetic variants with neutralizing antibody response to measles vaccine. Hum Genet 136, 421–435 (2017). https://doi.org/10.1007/s00439-017-1768-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-017-1768-9