Abstract
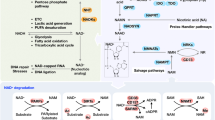
NAD+ (nicotinamide adenine dinucleotide) is an essential metabolite involved in a myriad of cellular processes. The NAD+ pool is maintained by three biosynthesis pathways, which are largely conserved from bacteria to human with some species-specific differences. Studying the regulation of NAD+ metabolism has been difficult due to the dynamic flexibility of NAD+ intermediates, the redundancy of biosynthesis pathways, and the complex interconnections among them. The budding yeast Saccharomyces cerevisiae provides an efficient genetic model for the isolation and study of factors that regulate specific NAD+ biosynthesis pathways. A recent study has uncovered a putative cross-regulation between the de novo NAD+ biosynthesis and copper homeostasis mediated by a copper-sensing transcription factor Mac1. Mac1 appears to work with the Hst1–Sum1–Rfm1 complex to repress the expression of de novo NAD+ biosynthesis genes. Here, we extend the discussions to include additional nutrient- and stress-sensing pathways that have been associated with the regulation of NAD+ homeostasis. NAD+ metabolism is an emerging therapeutic target for several human diseases. NAD+ preservation also helps ameliorate age-associated metabolic disorders. Recent findings in yeast contribute to the understanding of the molecular basis underlying the cross-regulation of NAD+ metabolism and other signaling pathways.


Similar content being viewed by others
References
Amaral M, Outeiro TF, Scrutton NS, Giorgini F (2013) The causative role and therapeutic potential of the kynurenine pathway in neurodegenerative disease. J Mol Med (Berlin, Germany) 91:705–713. https://doi.org/10.1007/s00109-013-1046-9
Anderson RM et al (2002) Manipulation of a nuclear NAD+ salvage pathway delays aging without altering steady-state NAD+ levels. J Biol Chem 277:18881–18890. https://doi.org/10.1074/jbc.m111773200
Anderson RM, Bitterman KJ, Wood JG, Medvedik O, Sinclair DA (2003) Nicotinamide and PNC1 govern lifespan extension by calorie restriction in Saccharomyces cerevisiae. Nature 423:181–185. https://doi.org/10.1038/nature01578
Bedalov A, Hirao M, Posakony J, Nelson M, Simon JA (2003) NAD+ -dependent deacetylase Hst1p controls biosynthesis and cellular NAD+ levels in Saccharomyces cerevisiae. Mol Cell Biol 23:7044–7054
Belenky P, Racette FG, Bogan KL, McClure JM, Smith JS, Brenner C (2007) Nicotinamide riboside promotes Sir2 silencing and extends lifespan via Nrk and Urh1/Pnp1/Meu1 pathways to NAD+. Cell 129:473–484. https://doi.org/10.1016/j.cell.2007.03.024
Belenky PA, Moga TG, Brenner C (2008) Saccharomyces cerevisiae YOR071C encodes the high affinity nicotinamide riboside transporter Nrt1. J Biol Chem 283:8075–8079. https://doi.org/10.1074/jbc.c800021200
Bieganowski P, Brenner C (2004) Discoveries of nicotinamide riboside as a nutrient and conserved NRK genes establish a Preiss-Handler independent route to NAD+ in fungi and humans. Cell 117:495–502
Bieganowski P, Pace HC, Brenner C (2003) Eukaryotic NAD synthetase Qns1 contains an essential, obligate intramolecular thiol glutamine amidotransferase domain related to nitrilase. J Biol Chem 278:33049–33055. https://doi.org/10.1074/jbc.m302257200
Bitterman KJ, Anderson RM, Cohen HY, Latorre-Esteves M, Sinclair DA (2002) Inhibition of silencing and accelerated aging by nicotinamide, a putative negative regulator of yeast sir2 and human SIRT1. J Biol Chem 277:45099–45107. https://doi.org/10.1074/jbc.m205670200
Bogan KL, Evans C, Belenky P, Song P, Burant CF, Kennedy R, Brenner C (2009) Identification of Isn1 and Sdt1 as glucose- and vitamin-regulated nicotinamide mononucleotide and nicotinic acid mononucleotide [corrected] 5’-nucleotidases responsible for production of nicotinamide riboside and nicotinic acid riboside. J Biol Chem 284:34861–34869. https://doi.org/10.1074/jbc.m109.056689
Brachmann CB, Sherman JM, Devine SE, Cameron EE, Pillus L, Boeke JD (1995) The SIR2 gene family, conserved from bacteria to humans, functions in silencing, cell cycle progression, and chromosome stability. Genes Dev 9:2888–2902
Braidy N, Grant R (2017) Kynurenine pathway metabolism and neuroinflammatory disease. Neural Regen Res 12:39–42. https://doi.org/10.4103/1673-5374.198971
Breda C et al (2016) Tryptophan-2,3-dioxygenase (TDO) inhibition ameliorates neurodegeneration by modulation of kynurenine pathway metabolites. Proc Natl Acad Sci USA 113:5435–5440. https://doi.org/10.1073/pnas.1604453113
Brown KD et al (2014) Activation of SIRT3 by the NAD(+) precursor nicotinamide riboside protects from noise-induced hearing loss. Cell Metab 20:1059–1068. https://doi.org/10.1016/j.cmet.2014.11.003
Canto C, Menzies KJ, Auwerx J (2015) NAD(+) metabolism and the control of energy homeostasis: a balancing act between mitochondria and the nucleus. Cell Metab 22:31–53. https://doi.org/10.1016/j.cmet.2015.05.023
Carroll AS, Bishop AC, DeRisi JL, Shokat KM, O’Shea EK (2001) Chemical inhibition of the Pho85 cyclin-dependent kinase reveals a role in the environmental stress response. Proc Natl Acad Sci USA 98:12578–12583. https://doi.org/10.1073/pnas.211195798
Chandler JL, Gholson RK (1972) De novo biosynthesis of nicotinamide adenine dinucleotide in Escherichia coli: excretion of quinolinic acid by mutants lacking quinolinate phosphoribosyl transferase. J Bacteriol 111:98–102
Chang KH, Cheng ML, Tang HY, Huang CY, Wu YR, Chen CM (2018) Alternations of metabolic profile and kynurenine metabolism in the plasma of Parkinson’s disease. Mol Neurobiol 55:6319–6328. https://doi.org/10.1007/s12035-017-0845-3
Chini CC, Tarrago MG, Chini EN (2016) NAD and the aging process: role in life, death and everything in between. Mol Cell Endocrinol. https://doi.org/10.1016/j.mce.2016.11.003
Croft T, James Theoga Raj C, Salemi M, Phinney BS, Lin SJ (2018) A functional link between NAD(+) homeostasis and N-terminal protein acetylation in Saccharomyces cerevisiae. J Biol Chem 293:2927–2938. https://doi.org/10.1074/jbc.m117.807214
Emanuelli M, Carnevali F, Lorenzi M, Raffaelli N, Amici A, Ruggieri S, Magni G (1999) Identification and characterization of YLR328W, the Saccharomyces cerevisiae structural gene encoding NMN adenylyltransferase. Expression and characterization of the recombinant enzyme. FEBS Lett 455:13–17
Frye RA (2000) Phylogenetic classification of prokaryotic and eukaryotic Sir2-like proteins. Biochem Biophys Res Commun 273:793–798
Garten A, Schuster S, Penke M, Gorski T, de Giorgis T, Kiess W (2015) Physiological and pathophysiological roles of NAMPT and NAD metabolism. Nat Rev Endocrinol 11:535–546. https://doi.org/10.1038/nrendo.2015.117
Ghislain M, Talla E, Francois JM (2002) Identification and functional analysis of the Saccharomyces cerevisiae nicotinamidase gene, PNC1. Yeast 19:215–224. https://doi.org/10.1002/yea.810
Graden JA, Winge DR (1997) Copper-mediated repression of the activation domain in the yeast Mac1p transcription factor. Proc Natl Acad Sci USA 94:5550–5555
Grose JH, Bergthorsson U, Roth JR (2005) Regulation of NAD synthesis by the trifunctional NadR protein of Salmonella enterica. J Bacteriol 187:2774–2782. https://doi.org/10.1128/JB.187.8.2774-2782.2005
Gross C, Kelleher M, Iyer VR, Brown PO, Winge DR (2000) Identification of the copper regulon in Saccharomyces cerevisiae by DNA microarrays. J Biol Chem 275:32310–32316. https://doi.org/10.1074/jbc.M005946200
Grozio A et al (2019) Slc12a8 is a nitotinamide mononucleotide transporter. Nat Metab 1:47–57. https://doi.org/10.1038/s42255-018-0009-4
Imai S, Guarente L (2014) NAD+ and sirtuins in aging and disease. Trends Cell Biol 24:464–471. https://doi.org/10.1016/j.tcb.2014.04.002
Jackson MD, Schmidt MT, Oppenheimer NJ, Denu JM (2003) Mechanism of nicotinamide inhibition and transglycosidation by Sir2 histone/protein deacetylases. J Biol Chem 278:50985–50998. https://doi.org/10.1074/jbc.m306552200
James Theoga Raj C, Croft T, Venkatakrishnan P, Groth B, Dhugga G, Cater T, Lin SJ (2019) The copper-sensing transcription factor Mac1, the histone deacetylase Hst1, and nicotinic acid regulate de novo NAD(+) biosynthesis in budding yeast. J Biol Chem. https://doi.org/10.1074/jbc.ra118.006987
Jensen LT, Posewitz MC, Srinivasan C, Winge DR (1998) Mapping of the DNA binding domain of the copper-responsive transcription factor Mac1 from Saccharomyces cerevisiae. J Biol Chem 273:23805–23811
Jungmann J, Reins HA, Lee J, Romeo A, Hassett R, Kosman D, Jentsch S (1993) MAC1, a nuclear regulatory protein related to Cu-dependent transcription factors is involved in Cu/Fe utilization and stress resistance in yeast. EMBO J 12:5051–5056
Kato M, Lin SJ (2014a) Regulation of NAD+ metabolism, signaling and compartmentalization in the yeast Saccharomyces cerevisiae. DNA Repair 23:49–58. https://doi.org/10.1016/j.dnarep.2014.07.009
Kato M, Lin SJ (2014b) YCL047C/POF1 is a novel nicotinamide mononucleotide adenylyltransferase (NMNAT) in Saccharomyces cerevisiae. J Biol Chem 289:15577–15587. https://doi.org/10.1074/jbc.m114.558643
Katsyuba E et al (2018) De novo NAD(+) synthesis enhances mitochondrial function and improves health. Nature 563:354–359. https://doi.org/10.1038/s41586-018-0645-6
Kulikova V et al (2015) Generation, release, and uptake of the NAD precursor nicotinic acid riboside by human cells. J Biol Chem 290:27124–27137. https://doi.org/10.1074/jbc.M115.664458
Lin JB et al (2016) NAMPT-mediated NAD(+) biosynthesis is essential for vision in mice. Cell Rep 17:69–85. https://doi.org/10.1016/j.celrep.2016.08.073
Liu HW et al (2018) Pharmacological bypass of NAD(+) salvage pathway protects neurons from chemotherapy-induced degeneration. Proc Natl Acad Sci USA 115:10654–10659. https://doi.org/10.1073/pnas.1809392115
Llorente B, Dujon B (2000) Transcriptional regulation of the Saccharomyces cerevisiae DAL5 gene family and identification of the high affinity nicotinic acid permease TNA1 (YGR260w). FEBS Lett 475:237–241
Lu SP, Lin SJ (2010) Regulation of yeast sirtuins by NAD(+) metabolism and calorie restriction. Biochim Biophys Acta 1804:1567–1575. https://doi.org/10.1016/j.bbapap.2009.09.030
Lu SP, Lin SJ (2011) Phosphate-responsive signaling pathway is a novel component of NAD+ metabolism in Saccharomyces cerevisiae. J Biol Chem 286:14271–14281. https://doi.org/10.1074/jbc.m110.217885
Lu SP, Kato M, Lin SJ (2009) Assimilation of endogenous nicotinamide riboside is essential for calorie restriction-mediated life span extension in Saccharomyces cerevisiae. J Biol Chem 284:17110–17119. https://doi.org/10.1074/jbc.M109.004010
Medvedik O, Lamming DW, Kim KD, Sinclair DA (2007) MSN2 and MSN4 link calorie restriction and TOR to sirtuin-mediated lifespan extension in Saccharomyces cerevisiae. PLoS Biol 5:e261. https://doi.org/10.1371/journal.pbio.0050261
Mole DJ et al (2016) Kynurenine-3-monooxygenase inhibition prevents multiple organ failure in rodent models of acute pancreatitis. Nat Med 22:202–209. https://doi.org/10.1038/nm.4020
Nikiforov A, Dolle C, Niere M, Ziegler M (2011) Pathways and subcellular compartmentation of NAD biosynthesis in human cells: from entry of extracellular precursors to mitochondrial NAD generation. J Biol Chem 286:21767–21778. https://doi.org/10.1074/jbc.M110.213298
Nikiforov A, Kulikova V, Ziegler M (2015) The human NAD metabolome: functions, metabolism and compartmentalization. Crit Rev Biochem Mol Biol 50:284–297. https://doi.org/10.3109/10409238.2015.1028612
Ohashi K, Kawai S, Murata K (2013) Secretion of quinolinic acid, an intermediate in the kynurenine pathway, for utilization in NAD+ biosynthesis in the yeast Saccharomyces cerevisiae. Eukaryot Cell 12:648–653. https://doi.org/10.1128/ec.00339-12
Ohashi K, Chaleckis R, Takaine M, Wheelock CE, Yoshida S (2017) Kynurenine aminotransferase activity of Aro8/Aro9 engage tryptophan degradation by producing kynurenic acid in Saccharomyces cerevisiae. Sci Rep 7:12180. https://doi.org/10.1038/s41598-017-12392-6
Panozzo C et al (2002) Aerobic and anaerobic NAD+ metabolism in Saccharomyces cerevisiae. FEBS Lett 517:97–102
Poyan Mehr A et al (2018) De novo NAD(+) biosynthetic impairment in acute kidney injury in humans. Nat Med 24:1351–1359. https://doi.org/10.1038/s41591-018-0138-z
Ratajczak J et al (2016) NRK1 controls nicotinamide mononucleotide and nicotinamide riboside metabolism in mammalian cells. Nat Commun 7:13103. https://doi.org/10.1038/ncomms13103
Revollo JR, Grimm AA, Imai S (2004) The NAD biosynthesis pathway mediated by nicotinamide phosphoribosyltransferase regulates Sir2 activity in mammalian cells. J Biol Chem 279:50754–50763. https://doi.org/10.1074/jbc.m408388200
Said HM, Nabokina SM, Balamurugan K, Mohammed ZM, Urbina C, Kashyap ML (2007) Mechanism of nicotinic acid transport in human liver cells: experiments with HepG2 cells and primary hepatocytes. Am J Physiol Cell Physiol 293:C1773–C1778. https://doi.org/10.1152/ajpcell.00409.2007
Schwarcz R, Bruno JP, Muchowski PJ, Wu HQ (2012) Kynurenines in the mammalian brain: when physiology meets pathology. Nat Rev Neurosci 13:465–477. https://doi.org/10.1038/nrn3257
Serpe M, Joshi A, Kosman DJ (1999) Structure-function analysis of the protein-binding domains of Mac1p, a copper-dependent transcriptional activator of copper uptake in Saccharomyces cerevisiae. J Biol Chem 274:29211–29219
Sporty J, Lin SJ, Kato M, Ognibene T, Stewart B, Turteltaub K, Bench G (2009) Quantitation of NAD+ biosynthesis from the salvage pathway in Saccharomyces cerevisiae. Yeast 26:363–369. https://doi.org/10.1002/yea.1671
Swinnen E et al (2006) Rim15 and the crossroads of nutrient signalling pathways in Saccharomyces cerevisiae. Cell Div 1:3. https://doi.org/10.1186/1747-1028-1-3
Tavares RG, Tasca CI, Santos CE, Alves LB, Porciuncula LO, Emanuelli T, Souza DO (2002) Quinolinic acid stimulates synaptosomal glutamate release and inhibits glutamate uptake into astrocytes. Neurochem Int 40:621–627
Trammell SA et al (2016) Nicotinamide riboside is uniquely and orally bioavailable in mice and humans. Nat Commun 7:12948. https://doi.org/10.1038/ncomms12948
Tsang F, Lin SJ (2015) Less is more: Nutrient limitation induces cross-talk of nutrient sensing pathways with NAD(+) homeostasis and contributes to longevity. Front Biol (Beijing) 10:333–357. https://doi.org/10.1007/s11515-015-1367-x
Tsang F, James C, Kato M, Myers V, Ilyas I, Tsang M, Lin SJ (2015) Reduced Ssy1-Ptr3-Ssy5 (SPS) signaling extends replicative life span by enhancing NAD+ homeostasis in Saccharomyces cerevisiae. J Biol Chem 290:12753–12764. https://doi.org/10.1074/jbc.m115.644534
Verdin E (2015) NAD(+) in aging, metabolism, and neurodegeneration. Science 350:1208–1213. https://doi.org/10.1126/science.aac4854
Wei M, Fabrizio P, Hu J, Ge H, Cheng C, Li L, Longo VD (2008) Life span extension by calorie restriction depends on Rim15 and transcription factors downstream of Ras/PKA, Tor, and Sch9. PLoS Genet 4:e13. https://doi.org/10.1371/journal.pgen.0040013
Williams PA et al (2017) Vitamin B3 modulates mitochondrial vulnerability and prevents glaucoma in aged mice. Science 355:756–760. https://doi.org/10.1126/science.aal0092
Yang Y, Sauve AA (2016) NAD+ metabolism: Bioenergetics, signaling and manipulation for therapy. Biochim Biophys Acta 1864:1787–1800. https://doi.org/10.1016/j.bbapap.2016.06.014
Yoshino J, Baur JA, Imai SI (2018) NAD(+) intermediates: the biology and therapeutic potential of NMN and NR. Cell Metab 27:513–528. https://doi.org/10.1016/j.cmet.2017.11.002
Zhang H et al (2016) NAD+ repletion improves mitochondrial and stem cell function and enhances life span in mice. Science 352:1436–1443
Zhu Z, Labbe S, Pena MM, Thiele DJ (1998) Copper differentially regulates the activity and degradation of yeast Mac1 transcription factor. J Biol Chem 273:1277–1280
Acknowledgements
This work is supported by NIGMS, National institute of Health, Grant GM102297. The authors declare that they have no conflicts of interest with the contents of the article.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kupiec.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
James Theoga Raj, C., Lin, SJ. Cross-talk in NAD+ metabolism: insights from Saccharomyces cerevisiae. Curr Genet 65, 1113–1119 (2019). https://doi.org/10.1007/s00294-019-00972-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-019-00972-0