Abstract
Recent research has shown that cell spreading is highly dependent on the contractility of its cytoskeleton and the mechanical properties of the environment it is located in. The dynamics of such process is critical for the development of tissue engineering strategy but is also a key player in wound contraction, tissue maintenance and angiogenesis. To better understand the underlying physics of such phenomena, the paper describes a mathematical formulation of cell spreading and contraction that couples the processes of stress fiber formation, protrusion growth through actin polymerization at the cell edge and dynamics of cross-membrane protein (integrins) enabling cell-substrate attachment. The evolving cell’s cytoskeleton is modeled as a mixture of fluid, proteins and filaments that can exchange mass and generate contraction. In particular, besides self-assembling into stress fibers, actin monomers able to polymerize into an actin meshwork at the cell’s boundary in order to push the membrane forward and generate protrusion. These processes are possible via the development of cell-substrate attachment complexes that arise from the mechano-sensitive equilibrium of membrane proteins, known as integrins. After deriving the governing equation driving the dynamics of cell evolution and spreading, we introduce a numerical solution based on the extended finite element method, combined with a level set formulation. Numerical simulations show that the proposed model is able to capture the dependency of cell spreading and contraction on substrate stiffness and chemistry. The very good agreement between model predictions and experimental observations suggests that mechanics plays a strong role into the coupled mechanisms of contraction, adhesion and spreading of adherent cells.












Similar content being viewed by others
References
Abraham VC, Krishnamurthi V, Taylor DL, Lanni F (1999) The actin-based nanomachine at the leading edge of migrating cells. Biophys J 77:1721–1732
Ateshian GA, Likhitpanichkul M, Hung CT (2006) A mixture theory analysis for passive transport in osmotic loading of cells. J Biomech 39:464–475
Ateshian GA (2007) On the theory of reactive mixtures for modeling biological growth. Biomech Model Mechanobiol 6(6):423–445
Baxter SC, Morales MO, Goldsmith EC (2008) Adaptive changes in cardiac fibroblast morphology and collagen organization as a result of mechanical environment. Cell Biochem Biophys 51:33–44
Bell GI (1978) Models for the specific adhesion of cells to cells. Science 200:618–627
Bell GI, Dembo M, Bongrand P (1984) Cell adhesion: competition between nonspecific repulsion and specific bonding. Biophys J 45:1051–1064
Belytschko T, Parimi Ch, Moes N, Sukumar N, Usui Sh (2003) Structured extended finite element methods for solids defined by implicit surfaces. Int J Numer Methods Eng 56(4):609–635
Bershadsky AD, Balaban NQ, Geiger B (2003) Adhesion dependent cell mechanosensitivity. Annu Rev Cell Dev Biol 19:677–695
Bischofs IB, Schwarz US (2003) Cell organization in soft media due to active mechanosensing. Proc Natl Acad Sci 100(16):9274–9279
Bottino D, Mogilner A, Roberts T, Stewart M, Oster G (2002) How nematode sperm crawl. J Cell Sci 115:367–384
Civelekoglu-Scholey G, Wayne Orr A, Novakd I, Meistera JJ, Schwartze MA, Mogilner A (2005) Model of coupled transient changes of rac, rho, adhesions and stress fibers alignment in endothelial cells responding to shear stress. J Theor Biol 48:569–585
Costa KD, Lee EJ, Holmes JW (2003) Creating alignement and anisotropy and engineering heart tissue: role of boundary conditions in a model three-dimensional culture system. Tissue Eng 9(4):567–577
Cramer LP (1997) Molecular mechanism of actin-dependent retrograde flow in lamellipodia of motile cells. Front Biosci 2:d26–270
Cramer LP, Mitchison TJ, Theriot JA (1994) Actin-dependent motile forces and cell motility. Curr Opin Cell Biol 6:82–86
Damien C, Thery M, Chu Y-S, Dufour S, Thiery J-P, Bornens M, Nassoy P, Mahadevan L (2007) The universal dynamics of cell spreading. Curr Biol 17:694–699
Deshpande VS, Mrksichb M, McMeekingc RM, Evans AG (2008) A bio-mechanical model for coupling cell contractility with focal adhesion formation. J Mech Phys Solids 56:1484–1510
Deshpande VS, McMeeking RM, Evans AG (2006) A bio-chemo-mechanical model for cell contractibility. Proc Natl Acad Sci 103(38):14015–14020
Dhote V, Vernerey FJ (2013) Mathematical model of the role of degradation on matrix development in hydrogel scaffolds (under review)
Dhote V, Skaalure S, Akalp U, Roberts J, Bryant SJ, Vernerey FJJ (2012) On the role of hydrogel structure and degradation in controlling the transport of cell-secreted matrix molecules for engineered cartilage. J Mech Behav Biomed Mater 12:S1751–6161
Deshpande VS, Mrksich M, McMeeking RM, Evans AG (2008) A bio-mechanical model for coupling cell contractility with focal adhesion formation. J Mech Phys Solids 56:1484–1510
DiMilla PA, Barbee K, Lauffenburger DA (1991) Mathematical model for the effects of adhesion and mechanics on cell migration speed. Biophys J 60:15–37
Dolbow J, Moes N, Belytschko T (2001) An extended finite element method for modeling crack growth with frictional contact. Comput Methods Appl Mech Eng 190(51–52):6825–6846
Duddu R, Bordas S, Chop D, Moran B (2008) A combined extended finite element and level set method for biofilm. IJNME 74:848–870
Farsad M, Vernerey F (2011) A multiphysics computational strategy for the modelling of cell-substrate interactions. Int J Numer Methods Eng. Page under review
Farsad M, Vernerey FJ, Park HS (2010) An extended finite element/level set method to study surface effects on the mechanical behavior and properties of nanomaterials. Int J Numer Methods Eng 84(12):1466–1489
Fernandez P, Bausch AR (2009) The compaction of gels by cells: a case of collective mechanical activity. Integr Biol 1:252–259
Foucard L, Vernerey FJ (2012) A thermodynamical model for stress-fiber organization in contractile cells. Appl Phys Lett 100:013702
Freyman T, Yannas I, Yokoo R, Gibson LJ (2001) Fibroblast contraction of a collagen-gag matrix. Biomaterials 22:2883–2891
Ghibaudo M, Saez A, Trichet L, Xayaphoummine A, Browaeys J, Silberzan P, Buguinb A, Ladoux B (2008) Traction forces and rigidity sensing regulate cell functions. Soft Mater 4:1836–1843
Gov NS, Gopinathan A (2006) Dynamics of membranes driven by actin polymerization. Biophys J 90:454–469
Greenwald E, Vernerey FJ, Bryant S (2012) Triphasic mixture model of cell-mediated enzymatic degradation of hydrogels. Comput Methods Biomech Biomed Eng 15(11):1197–1210
Grinnel F, Lamke CR (1984) Reorganization of hydrated collagen lattices by human skin fibroblasts. J Cell Sci 66:51–63
Guilak F, Mow VC (2000) The mechanical environment of the chondrocyte: a biphasic finite element model of cell-matrix interactions in articular cartilage. J Biomech 33:1663–1673
Guo WH, Frey MT, Burnham NA, Wang YL (2006) Substrate rigidity regulates the formation and maintenance of tissues. Biophys J 90:2213–2220
Guolla L, Bertrand M, Haase K, Pelling AE (2011) Force transduction and strain dynamics in actin stress fibres in response to nanonewton forces. J Cell Sci 125(3):603–613
Herant M, Dembo M (2010) Form and function in cell motility: from fibroblasts to keratocytes. Biophys J 98:1408–1417
Hettich T, Hund A, Ramm E (2008) Modeling of failure in composites by x-fem and level sets within a multiscale framework. Comput Methods Appl Mech Eng 197:414–424
Hill AV (1938) The heat of shortening and the dynamic constant of muscles. Proc R Soc B 126:136–195
Hill TL (1981) Microfilament or micrtubule assembly or disassembly against a force. Proc Natl Acad Sci USA 78(9):5613–5617
Hochmuth RM, Mohandas N, Blachshear PL (1973) Measurment of the elastic modulus for red cell membrane using a fluid mechanical technique. Biophys J 13:747–762
Hodge N, Papadopoulos P (2012) Continuum modeling and numerical simulation of cell motility. J Math Biol 64(7):1253–1279
Julicher F, Kruse K, Prost J, Joanny JF (2007) Active behavior of the cytoskeleton. Phys Rep 449:3–28
Kawabata K, Sado Y, Nagayama M, Nitta T, Nemoto K, Koyama Y, Haga H (2003) Visualization of dynamic organization of cytoskeleton gels in living cells by hybrid-SPM. Chin J Polym Sci 2:169–174
Koestler SA, Rottner K, Lai F, Block J, Vinzenz M, Small JV (2009) F- and g-actin concentrations in lamellipodia of moving cells. Plos One 4(3):e4810. doi: 10.1371/journal.pone.0004810
Kozlov MM, Bershadsky AD (2004) Processive capping by formin suggests a forcedriven mechanism of actin polymerization. J Cell Biol 167:1011–1017
Kumar S, Maxwell IZ, Heisterkamp A, Polte TR, Lele TP, Salanga M, Mazur E, Ingber DE (2006) Viscoelastic retraction of single living stress fibers and its impact on cell shape, cytoskeletal organization, and extracellular matrix mechanics. Biophys J 85:3762–3773
Lauffenburger DA, Linderman JJ (1993) Receptors models for binding, trafficking, and signaling. Oxford University Press, New York
Levental I, Georges PC, Janmey PA (2006) Soft biological materials and their impact on cell function. Soft Matter 2:1–9
Li Y, Xu G-K, Li B, Feng X-Q (2010) A molecular mechanisms-based biophysical model for two-phase cell spreading. Appl Phys Lett 96(043703):1–3
Lo CM, Wang HB, Dembo M, Wang YL (2000) Cell movement is guided by the rigidity of the substrate. Biophys J 79:144–152
Moes N, Cloirec M, Cartraud P, Remacle JF (2003) A computational approach to handle complex microstructure geometries. Comput Methods Appl Mech Eng 192:3163–3177
Mogilner A, Rubinstein B (2005) The physics of filopodial protrusion. Biophys J 89:782–795
Mohammadi S (2008) Extended finite element method. Blackwell, Oxford
Nicolas A, Besser A, Safran SA (2008) Dynamics of cellular focal adhesions on deformable substrates: consequences for cell force microscopy. Biophys J 95:527–539
Novak IL, Slepchenko BM, Mogilner A (2008) Quantitative analysis of g-actin transport in motile cells. Biophys J 95:1627–1638
Oster GF, Perelson AS (1985) Cell spreading and motility: a model lamellipod. J Math Biol 21:383–388
Parker KK, Brock AL, Brangwynne C, Mannix RJ, Wang N, Ostuni E, Geisse NA, Adams JC, Whitesides GM, Ingber DE (2002) Directional control of lamellipodia extension by constraining cell shape and orienting cell tractional forces. FASEB J 16:1195–1204
Parker KK, Ingber DE (2007) Extracellular matrix, mechanotransduction and structural hierarchies in heart tissue engineering. Philos Trans R Soc B 362:1267–1279
Pelham RJ, Wang YL (1997) Cell locomotion and focal adhesions are regulated by substrate flexibility. Cell Biol 94:13661–13665
Peskin CS, Odell GM, Oster GF (1993) Cellular motions and thermal fluctuations: the brownian ratchet. Biophys J 65:316–324
Petrol WM, Ma L, Jester JV (2003) Direct correlation of collagen matrix deformation with focal adhesion dynamics in living corneal fibroblasts. J Cell Sci 56:1484–1510
Pollard TD, Borisy GG (2003) Cellular motility driven by assembly and disassembly of actin filaments. Cell 112:453–465
Rafelski SM, Theriot JA (2004) Crawling toward a unified model of cell motility: spatial and temporal regulation of actin dynamics. Annual Rev Biochem 73:209–239
Reinhart-King CA, Dembo M, Hammer DA (2005) The dynamics and mechanics of endothelial cell spreading. Biophys J 89:676–689
Rubinstein B, Jacobson K, Mogilner A (2005) Multiscale two-dimensional modeling of a motile simple-shaped cell. Multiscale Model Simul 3(2):413–439
Schwarz US, Bischofs IB (2005) Physical determinants of cell organization in soft media. Med Eng Phys 27:763–772
Shemesh T, Bershadsky AD, Kozlov MM (2005) Force driven polymerization in cells: actin filaments and focal adhesions. J Phys Condens Matter 17:S3913–S3928
Shemesh T, Geiger B, Bershadsky A, Kozlov MM (2005) Focal adhesions as mechanosensors: a physical mechanism. Proc Natl Acad Sci 102:12383–12388
Solon J, Levental I, Sengupta K, Georges PC, Janmey PA (2007) Fibroblast adaptation and stiffness matching to soft elastic substrates. Biophys J 93(12):4453–4461
Stamenovic D, Lazopoulos KA, Pirentis A, Suki BE (2009) Mechanical stability determines stress fiber and focal adhesion orientation. Cell Mol Bioeng 2(4):475–485
Tan JL, Tien J, Pirone DM, Gray DS, Bhadriraju K, Chen CS (2003) Cells lying on a bed of microneedles: an approach to isolate mechanical force. Proc Natl Acad Sci 100(4):1484–1489
Tsuda Y, Yasutake H, Ishijima A, Yanagida T (1996) Torsional rigidity of single actin filaments and actin–actin bond breaking force under torsion measured directly by in vitro micromanipulation. Proc Natl Acad Sci 93:12937–12942
Vallotton P, Danuser G, Bohnet S, Meister J-J, Verkhovsky AB (2005) Tracking retrograde flow in keratocytes: news from the front. Mol Biol Cell 16:1223–1231
Vernerey FJ (2011) A theoretical treatment on the mechanics of interfaces in deformable porous media. Int J Solids Struct 48(22–23):1267–1279
Vernerey FJ (2012) On the application of multiphasic theories to the problem of cell-substrate mechanical interactions. Adv Cell Mech 189–224
Vernerey FJ, Farsad M (2011) An Eulerian/XFEM formulation for the large deformation of cortical cell membrane. Comput Methods Biomech Biomed Eng 14(5):433–445
Vernerey FJ, Farsad M (2011) A constrained mixture approach to mechanosensing and force generation in contractile cells. J Mech Behav Biomed Mater 4:1683–1699
Wakatsuki T, Wysolmerski RB, Elson EL (2003) Mechanics of cell spreading: role of myosin II. J Cell Sci 116:1617–1625
Walcott S, Sun SX (2009) A mechanical model of actin stress fiber formation and substrate elasticity sensing in adherent cells. Proc Natl Acad Sci 107(17):7757–7762
Wang HB, Dembo M, Wang YL (2000) Substrate flexibility regulates growth and apoptosis of normal but not transformed cells. Am J Physiol Cell Physiol 279:C1345–C1350
White GE, Fujiwara K (1986) Expression and intracellular distribution of stress fibers in aortic endothelium. J Cell Biol 103:63–70
Xiong Y, Rangamani P, Fardin M-A, Lipshtat A, Dubin-Thaler B, Rossier O, Sheetz MP, Iyengar R (2010) Mechanisms controlling cell size and shape during isotropic cell spreading. Biophys J 98:2136–2146
Yeung T, Georges PC, Flanagan LA, Marg B, Ortiz M, Funaki M, Zahir N, Ming W, Weaver V, Janmey PA (2005) Effects of substrate stiffness on cell morphology, cytoskeletal structure, and adhesion. Cell Motil Cytoskelet 60:24–34
Acknowledgments
FJV gratefully acknowledges the University of Colorado CRCW Seed Grant and NIH Grant Number 1R21AR061011 in support of this research. We would also like to acknowledge the reviewers for contributing to this work via their insightful comments and suggestions.
Author information
Authors and Affiliations
Corresponding author
Appendix: A
Appendix: A
1.1 A.1 Plane-stress conditions elasticity equations for cell
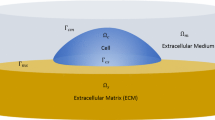
In this paper, it is assumed that the cell’s geometry can be described as a very thin plate such that two-dimensional plane-stress conditions can be applied. This implies that the total stress \(\mathbf T\) verifies \(T_{xz}=T_{yz}=0,T_{zz}=0\). Since stress fibers do not exert forces in the z-direction, the last equation can be written \(T^{c}_{zz} - p=0\), where \(p\) is the cytosol pressure. Writing the three-dimensional stress-strain relation for the actin filament network as:
and enforcing the fact that \(T^c_{zz}=p\), we obtain the following expression for \(\varepsilon _{zz}\):
where \(E\) and \(\nu \) are the Young’s modulus and Poisson’s ratio of the actin filament network. Their relationship with Lame’s constants appearing in (70) is given below:
We now wish to express the divergence \(\nabla \cdot \mathbf{v}^c\) of the velocity field \(\mathbf{v}^c\) for substitution in the equations of mass balance (4–6). For a three dimensional problem, the divergence reads \(\nabla \cdot \mathbf{v}^c={\dot{\varepsilon }}_{xx}+{\dot{\varepsilon }}_{yy} +{\dot{\varepsilon }}_{zz}\) where a superimposed dot is used to denote a time derivative. Using the fact that:
from (71), we can rewrite:
where \(\nabla ^{(2)}\cdot \mathbf{v}=\partial v_x/\partial x+ \partial v_y/\partial y={\dot{\varepsilon }}_{xx}+ {\dot{\varepsilon }}_{yy}\) represents the divergence of the velocity \(\mathbf{v}\) in the two-dimensional plane \(xy\) while constants \(\alpha \) and \(\beta \) can be found to be:
In the particular case of axisymmetric conditions, (74) becomes:
This result is used to obtain Eqs. (7) and (8) in the present study.
1.2 A.2 Stiffness and damping matrices
It can be shown, through a standard finite-element procedure, that the force vector, damping matrix and stiffness matrix appearing in Eq. (63) take the following form:
where the superscript \(e\) denotes quantities associated with individual finite element \(e\) and the symbol \({\fancyscript{A}}\) denotes the assembly operator. The final force vector, damping matrix, and stiffness matrix finally take the forms:
and
where the superscripts \(ij\) is used to designate a matrix representing the interactions between component \(i\) and component \(j\).
Rights and permissions
About this article
Cite this article
Vernerey, F.J., Farsad, M. A mathematical model of the coupled mechanisms of cell adhesion, contraction and spreading. J. Math. Biol. 68, 989–1022 (2014). https://doi.org/10.1007/s00285-013-0656-8
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00285-013-0656-8