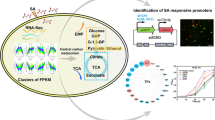
Abstract
Kasugamycin, produced by Streptomyces kasugaensis and Streptomyces microaureus, is an important amino-glycoside family antibiotic and widely used for veterinary and agricultural applications. In the left flanking region of the previously reported kasugamycin gene cluster, four additional genes (two-component system kasW and kasX, MerR-family kasV, and isoprenylcysteine carboxyl methyltransferase kasS) were identified both in the low-yielding S. kasugaensis BCRC12349 and high-yielding S. microaureus XM301. Deletion of regulatory gene kasT abolished kasugamycin production, and its overexpression in BCRC12349 resulted in an increased titer by 186 %. Deletion of kasW, kasX, kasV, and kasS improved kasugamycin production by 12, 19, 194, and 22 %, respectively. qRT-PCR analysis demonstrated that the transcription of kas genes was significantly increased in all the four mutants. Similar gene inactivation was performed in the high-yielding strain S. microaureus XM301. As expected, the deletion of kasW/X resulted in a 58 % increase of the yield from 6 to 9.5 g/L. However, the deletion of kasV and over-expression of kasT had no obvious effect, and the disruption of kasS surprisingly decreased kasugamycin production. In addition, trans-complementation of the kasS mutant with a TTA codon-mutated kasS increased the kasugamycin yield by 20 %. A much higher transcription of kas genes was detected in the high-yielding XM301 than in the low-yielding BCRC12349, which may partially account for the discrepancy of gene inactivation effects between them. Our work not only generated engineered strains with improved kasugamycin yield, but also pointed out that different strategies on manipulating regulatory-related genes should be considered for low-yielding or high-yielding strains.





Similar content being viewed by others
References
Baron RA, Casey PJ (2004) Analysis of the kinetic mechanism of recombinant human isoprenylcysteine carboxylmethyltransferase (ICMT). BMC Biochem 5:19
Berdy J (2005) Bioactive microbial metabolites. J Antibiot 58(1):1–26
Bibb MJ (2005) Regulation of secondary metabolism in Streptomycetes. Curr Opin Microbiol 8(2):208–15
Brown NL, Stoyanov JV, Kidd SP, Hobman JL (2003) The MerR family of transcriptional regulators. FEMS Microbiol Lett 27(2–3):145–163
Chater KF (2006) Streptomyces inside-out: a new perspective on the bacteria that provide us with antibiotics. Philos T R Soc B Biological Sci 361(1469):761–8
Chater KF, Chandra G (2008) The use of the rare UUA codon to define "expression space" for genes involved in secondary metabolism, development and environmental adaptation in Streptomyces. J Microbiol 46(1):1–11
Cheng L, Chen W, Zhai L, Xu D, Huang T, Lin S, Zhou X, Deng Z (2011) Identification of the gene cluster involved in muraymycin biosynthesis from Streptomyces sp. NRRL 30471. Mol BioSyst 7(3):920–7
Dong L, Nakashima N, Tamura N, Tamura T (2004) Isolation and characterization of the Rhodococcus opacus thiostrepton-inducible genes tipAL and tipAS: application for recombinant protein expression in Rhodococcus. FEMS Microbiol Lett 237(1):35–40
Flatt PM, Mahmud T (2007) Biosynthesis of aminocyclitol-aminoglycoside antibiotics and related compounds. Nat Prod Rep 24(2):358–92
Gust B, Challis GL, Fowler K, Kieser T, Chater KF (2003) PCR-targeted Streptomyces gene replacement identifies a protein domain needed for biosynthesis of the sesquiterpene soil odor geosmin. Proc Natl Acad Sci U S A 100(4):1541–6
Higo A, Horinouchi S, Ohnishi Y (2011) Strict regulation of morphological differentiation and secondary metabolism by a positive feedback loop between two global regulators AdpA and BldA in Streptomyces griseus. Mol Microbiol 81(6):1607–22
Hotta K, Ogata T, Ishikawa J, Okanishi M, Mizuno S, Morioka M, Naganawa H, Okami Y (1996) Mechanism of multiple aminoglycoside resistance of kasugamycin-producing Streptomyces kasugaensis MB273: involvement of two types of acetyltransferases in resistance to astromicin group antibiotics. J Antibiot 49(7):682–8
Huang J, Shi J, Molle V, Sohlberg B, Weaver D, Bibb MJ, Karoonuthaisiri N, Lih CJ, Kao CM, Buttner MJ, Cohen SN (2005) Cross-regulation among disparate antibiotic biosynthetic pathways of Streptomyces coelicolor. Mol Microbiol 58:1276–87
Hwang KS, Kim HU, Charusanti P, Palsson BO, Lee SY (2014) Systems biology and biotechnology of Streptomyces species for the production of secondary metabolites. Biotechnol Adv 32(2):255–68
Ikeno S, Aoki D, Hamada M, Hori M, Tsuchiya KS (2006) DNA sequencing and transcriptional analysis of the kasugamycin biosynthetic gene cluster from Streptomyces kasugaensis M338-M1. J Antibiot 59(1):18–28
Ikeno S, Aoki D, Sato K, Hamada M, Hori M, Tsuchiya KS (2002) kasT gene of Streptomyces kasugaensis M338-M1 encodes a DNA-binding protein which binds to intergenic region of kasU-kasJ in the kasugamycin biosynthesis gene cluster. J Antibiot 55(12):1053–62
Ikeno S, Higashide K, Kinoshita N, Hamada M, Hori M (1998) A 7.6-kb DNA region from Streptomyces kasugaensis M338-M1 includes some genes responsible for kasugamycin biosynthesis. J Antibiot 51:341–52
Ikeno S, Ohishi Y, Kinoshita N, Hamada M, Tsuchiya KS, Hori M (2000) ABC transporter genes, kasKLM, responsible for self-resistance of a kasugamycin producer strain. J Antibiot 53:373–84
Kaberdina AC, Szaflarski W, Nierhaus KH, Moll I (2009) An unexpected type of ribosomes induced by kasugamycin: a look into ancestral times of protein synthesis. Mol Cell 33(2):227–36
Kojima I, Kasuga K, Kobayashi M, Fukasawa A, Mizuno S, Arisawa A, Akagawa H (2002) The rpoZ gene, encoding the RNA polymerase omega subunit, is required for antibiotic production and morphological differentiation in Streptomyces kasugaensis. J Bacteriol 184(23):6417–23
Komatsu M, Komatsu K, Koiwai H, Yamada Y, Kozone I, Izumikawa M, Hashimoto J, Takagi M, Omura S, Shin-ya K, Cane DE, Ikeda H (2013) Engineered Streptomyces avermitilis host for heterologous expression of biosynthetic gene cluster for secondary metabolites. ACS Synth Biol 2(7):384–96
Leung KF, Baron R, Ali BR, Magee AI, Seabra MC (2007) Rab GTPases containing a CAAX motif are processed post-geranylgeranylation by proteolysis and methylation. J Biol Chem 282(2):1487–97
Liu G, Chater KF, Chandra G, Niu G, Tan H (2013) Molecular regulation of antibiotic biosynthesis in Streptomyces. Microbiol Mol Biol Rev 77:112–43
Martin JF, Liras P (2010) Engineering of regulatory cascades and networks controlling antibiotic biosynthesis in Streptomyces. Curr Opin Microbiol 13(3):263–73
Olano C, Lombo F, Mendez C, Salas JA (2008) Improving production of bioactive secondary metabolites in actinomycetes by metabolic engineering. Metab Eng 10(5):281–92
Paget MS, Chamberlin L, Atrih A, Foster SJ, Buttner MJ (1999) Evidence that the extracytoplasmic function sigma factor σE is required for normal cell wall structure in Streptomyces coelicolor A3(2). J Bacteriol 181(1):204–11
Qiu J, Zhuo Y, Zhu D, Zhou X, Zhang L, Bai L, Deng Z (2011) Overexpression of the ABC transporter AvtAB increases avermectin production in Streptomyces avermitilis. Appl Microbiol Biotechnol 92(2):337–45
Santos BF, Rodriguez GA, Sola LA, Martin JF (2009) Cross-talk between two global regulators in Streptomyces: PhoP and AfsR interact in the control of afsS, pstS and phoRP transcription. Mol Microbiol 72(1):53–68
Sola LA, Moura RS, Martin JF (2003) The two-component PhoR-PhoP system controls both primary metabolism and secondary metabolite biosynthesis in Streptomyces lividans. Proc Natl Acad Sci U S A 100:6133–8
Sola LA, Rodriguez GA, Amin R, Wohlleben W, Martin JF (2013) Competition between the GlnR and PhoP regulators for the glnA and amtB promoters in Streptomyces coelicolor. Nucleic Acids Res 41:1767–82
Tomono A, Tsai Y, Yamazaki H, Ohnishi Y, Horinouchi S (2005) Transcriptional control by A-factor of strR, the pathway-specific transcriptional activator for streptomycin biosynthesis in Streptomyces griseus. J Bacteriol 187(16):5595–604
Wang R, Mast Y, Wang J, Zhang W, Zhao G, Wohlleben W, Lu Y, Jiang W (2013) Identification of two-component system AfsQ1/Q2 regulon and its cross-regulation with GlnR in Streptomyces coelicolor. Mol Microbiol 87(1):30–48
Wilkinson CJ, Hughes T, Martin CJ, Bohm I, Mironenko T, Deacon M, Wheatcroft M, Wirtz G, Staunton J, Leadlay PF (2002) Increasing the efficiency of heterologous promoters in actinomycetes. J Mol Microbiol Biotechnol 4(4):417–426
Yang J, Kulkarni K, Manolaridis I, Zhang Z, Dodd RB, Mas DC, Barford D (2011) Mechanism of isoprenylcysteine carboxyl methylation from the crystal structure of the integral membrane methyltransferase ICMT. Mol Cell 44(6):997–1004
Yu Z, Zhu H, Dang F, Zhang W, Qin Z, Yang S, Tan H, Lu Y, Jiang W (2012) Differential regulation of antibiotic biosynthesis by DraR-K, a novel two-component system in Streptomyces coelicolor. Mol Microbiol 85(3):535–56
Zhao Y, Xiang S, Dai X, Yang K (2013) A simplified diphenylamine colorimetric method for growth quantification. Appl Microbiol Biotechnol 97:5069–77
Acknowledgments
This work was supported by grants from the Ministry of Science and Technology of the People’s Republic of China (nos. 2012AA02A706, 2012AA022107, and 2012CB721005), the National Natural Science Foundation of China (nNo. 31470157), and the Program of University of Michigan – Shanghai Jiao Tong University Collaboration on Biomedical Technology.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
Chenchen Zhu declares that she has no conflict of interest. Qianjin Kang declares that he has no conflict of interest. Linquan Bai declares that he has no conflict of interest. Lin Cheng declares that he has no conflict of interest. Zixin Deng declares that he has no conflict of interest.
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 620 kb)
Rights and permissions
About this article
Cite this article
Zhu, C., Kang, Q., Bai, L. et al. Identification and engineering of regulation-related genes toward improved kasugamycin production. Appl Microbiol Biotechnol 100, 1811–1821 (2016). https://doi.org/10.1007/s00253-015-7082-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-015-7082-3