Abstract
Microbial communities are comprised of complex assemblages of highly interactive taxa. We employed network analyses to identify and describe microbial interactions and co-occurrence patterns between microbial eukaryotes and bacteria at two locations within a low salinity (0.5–3.5 ppt) lake over an annual cycle. We previously documented that the microbial diversity and community composition within Lake Texoma, southwest USA, were significantly affected by both seasonal forces and a site-specific bloom of the harmful alga, Prymnesium parvum. We used network analyses to answer ecological questions involving both the bacterial and microbial eukaryotic datasets and to infer ecological relationships within the microbial communities. Patterns of connectivity at both locations reflected the seasonality of the lake including a large rain disturbance in May, while a comparison of the communities between locations revealed a localized response to the algal bloom. A network built from shared nodes (microbial operational taxonomic units and environmental variables) and correlations identified conserved associations at both locations within the lake. Using network analyses, we were able to detect disturbance events, characterize the ecological extent of a harmful algal bloom, and infer ecological relationships not apparent from diversity statistics alone.




Similar content being viewed by others
References
Fuhrman JA, Caron DA (2015) Heterotrophic planktonic microbes: virus, bacteria, archaea, and protozoa. In: Yates, MV, Nakatsu, CH, Miller, RV, Pillai, SD (eds.) Manual of environmental microbiology, fourth edition. American Society of Microbiology, pp. 4.2.2–1 - 4.2.2–34
Worden AZ, Follows MJ, Giovannoni SJ, Wilken S, Zimmerman AE, Keeling PJ (2015) Rethinking the marine carbon cycle: factoring in the multifarious lifestyles of microbes. Science 347. https://doi.org/10.1126/science.1257594
Cole JJ (1982) Interactions between bacteria and algae in aquatic ecosystems. Ann Rev Ecol Syst 13:291–314
Sherr EB, Sherr BF (2002) Significance of predation by protists in aquatic microbial food webs. Antonie Van Leeuwenhoek 81:293–308. https://doi.org/10.1023/a:1020591307260
Ibelings BW, De Bruin A, Kagami M, Rijkeboer M, Brehm M, Donk EV (2004) Host parasite interactions between freshwater phytoplankton and chytrid fungi (Chytridiomycota). J. Phycol. 40:437–453. https://doi.org/10.1111/j.1529-8817.2004.03117.x
Gilbert JA, Field D, Swift P, Newbold L, Oliver A, Smyth T, Somerfield PJ, Huse S, Joint I (2009) The seasonal structure of microbial communities in the western English Channel. Environ. Microbiol. 11:3132–3139. https://doi.org/10.1111/j.1462-2920.2009.02017.x
Gilbert JA, Steele JA, Caporaso JG, Steinbruck L, Reeder J, Temperton B, Huse S, McHardy AC, Knight R, Joint I, Somerfield P, Fuhrman JA, Field D (2012) Defining seasonal marine microbial community dynamics. ISME J 6:298–308. https://doi.org/10.1038/ismej.2011.107
Jones SE, Chiu CC, Kratz TK, Wu JT, Shadeand A, Mcmahon KD (2008) Typhoons initiate predictable change in aquatic bacterial communities. Limnol. Oceanogr. 53:1319–1326
Vigil P, Countway PD, Rose JM, Gobler CJ, Lonsdale DJ, Caron DA (2009) Rapid shifts in dominant taxa among microbial eukaryotes in estuarine ecosystems. Aq Microb Ecol 54:83–100
Kim DY, Countway PD, Gast RJ, Caron DA (2011) Rapid shifts in the structure and composition of a protistan assemblage during bottle incubations affect estimates of total protistan species richness. Microb. Ecol. 62:383–398. https://doi.org/10.1007/s00248-011-9816-9
Shade A, Read JS, Youngblut ND, Fierer N, Knight R, Kratz TK, Lottig NR, Roden EE, Stanley EH, Stombaugh J, Whitaker RJ, CH W, McMahon KD (2012) Lake microbial communities are resilient after a whole-ecosystem disturbance. ISME J 6:2153–2167
Ings TC, Montoya JM, Bascompte J, Blüthgen N, Brown L, Dormann CF, Edwards F, Figueroa D, Jacob U, Jones JI, Lauridsen RB, Ledger ME, Lewis HM, Olesen JM, Van Veen FJF, Warren PH, Woodward G (2009) Review: ecological networks—beyond food webs. J Animal Ecol 78:253–269. https://doi.org/10.1111/j.1365-2656.2008.01460.x
Poulin R (2010) Network analysis shining light on parasite ecology and diversity. Trends Parasitol. 26:492–498. https://doi.org/10.1016/j.pt.2010.05.008
Proulx SR, Promislow DE, Phillips PC (2005) Network thinking in ecology and evolution. Trends Ecol. Evol. 20:345–353. https://doi.org/10.1016/j.tree.2005.04.004
Fuhrman JA, Cram JA, Needham DM (2015) Marine microbial community dynamics and their ecological interpretation. Nat Rev Micro 13:133–146. https://doi.org/10.1038/nrmicro3417
Williams RJ, Howe A, Hofmockel KS (2014) Demonstrating microbial co-occurrence pattern analyses within and between ecosystems. Frontiers Microbiol 5:358. https://doi.org/10.3389/fmicb.2014.00358
Eiler A, Heinrich F, Bertilsson S (2012) Coherent dynamics and association networks among lake bacterioplankton taxa. ISME J 6:330–342. https://doi.org/10.1038/ismej.2011.113
Milici M, Deng Z-L, Tomasch J, Decelle J, Wos-Oxley ML, Wang H, Jáuregui R, Plumeier I, Giebel H-A, Badewien TH, Wurst M, Pieper DH, Simon M, Wagner-Döbler I (2016) Co-occurrence analysis of microbial taxa in the Atlantic Ocean reveals high connectivity in the free-living bacterioplankton. Frontiers Microbiol 7:649. https://doi.org/10.3389/fmicb.2016.00649
Steele JA, Countway PD, Xia L, Vigil PD, Beman JM, Kim DY, Chow C-ET, Sachdeva R, Jones AC, Schwalbach MS, Rose JM, Hewson I, Patel A, Sun F, Caron DA, Fuhrman JA (2011) Marine bacterial, archaeal and protistan association networks reveal ecological linkages. ISME J 5:1414–1425
Barberan A, Bates ST, Casamayor EO, Fierer N (2012) Using network analysis to explore co-occurrence patterns in soil microbial communities. ISME J 6:343–351. https://doi.org/10.1038/ismej.2011.119
de Menezes AB, Prendergast-Miller MT, Richardson AE, Toscas P, Farrell M, Macdonald LM, Baker G, Wark T, Thrall PH (2015) Network analysis reveals that bacteria and fungi form modules that correlate independently with soil parameters. Environ. Microbiol. 17:2677–2689. https://doi.org/10.1111/1462-2920.12559
Lupatini M, Suleiman A, Jacques R, Antoniolli Z, Ferreira A, Kuramae EE, Roesch L (2014) Network topology reveal high connectance levels and few key microbial genera within soils. Frontiers Environ Sci 2. https://doi.org/10.3389/fenvs.2014.00010
Faust K, Sathirapongsasuti JF, Izard J, Segata N, Gevers D, Raes J, Huttenhower C (2012) Microbial co-occurrence relationships in the human microbiome. PLoS Comp Biol 8:e1002606. https://doi.org/10.1371/journal.pcbi.1002606
Guidi L, Chaffron S, Bittner L, Eveillard D, Larhlimi A, Roux S, Darzi Y, Audic S, Berline L, Brum J, Coelho LP, Espinoza JCI, Malviya S, Sunagawa S, Dimier C, Kandels-Lewis S, Picheral M, Poulain J, Searson S, Tara Oceans Consortium C, Stemmann L, Not F, Hingamp P, Speich S, Follows M, Karp-Boss L, Boss E, Ogata H, Pesant S, Weissenbach J, Wincker P, Acinas SG, Bork P, de Vargas C, Iudicone D, Sullivan MB, Raes J, Karsenti E, Bowler C, Gorsky G (2016) Plankton networks driving carbon export in the oligotrophic ocean. Nature 532:465–470. https://doi.org/10.1038/nature16942
Hambright KD, Zamor RM, Easton JD, Glenn KL, Remmel EJ, Easton AC (2010) Temporal and spatial variability of an invasive toxigenic protist in a North American subtropical reservoir. Harmful Algae 9:568–577. https://doi.org/10.1016/j.hal.2010.04.006
Evardsen B, Imai I (2006) The ecology of harmful flagellates within Prymnesiophyceae and Raphidophyceae. In: Granéli E, Turner J (eds) Ecology of harmful algae. Springer-Verlag, Berlin, pp. 67–79
Henrikson JC, Gharfeh MS, Easton AC, Easton JD, Glenn KL, Shadfan M, Mooberry SL, Hambright KD, Cichewicz RH (2010) Reassessing the ichthyotoxin profile of cultured Prymnesium parvum (golden algae) and comparing it to samples collected from recent freshwater bloom and fish kill events in North America. Toxicon 55:1396–1404
Igarashi T, Satake M, Yasumoto T (1999) Structures and partial stereochemical assignments for prymnesin-1 and prymnesin-2: potent hemolytic and ichthyotoxic glycosides isolated from the red ride alga Prymnesium parvum. J. Am. Chem. Soc. 121:8499–8511. https://doi.org/10.1021/ja991740e
Tillman U (1998) Phagotrophy by a plastidic haptophyte, Prymnesium patelliferum. Aq Microb Ecol 14:155–160
Remmel EJ, Hambright KD (2012) Toxin-assisted micropredation: experimental evidence shows that contact micropredation rather than exotoxicity is the role of Prymnesium toxins. Ecol. Lett. 15:126–132. https://doi.org/10.1111/j.1461-0248.2011.01718.x
Fistarol GO, Legrand C, Graneli E (2003) Allelopathic effect of Prymnesium parvum on a natural plankton community. Mar. Ecol. Prog. Ser. 255:115–125
Martin-Cereceda M, Novarino G, Young JR (2003) Grazing by Prymnesium parvum on small planktonic diatoms. Aq Microb Ecol 33:191–199. https://doi.org/10.3354/ame033191
Skovgaard A, Hansen PJ (2003) Food uptake in the harmful alga Prymnesium parvum mediated by excreted toxins. Limnol. Oceanogr. 48:1161–1166
Nejstgaard JC, Solberg PT (1996) Repression of copepod feeding and fecundity by the toxic haptophyte Prymnesium patelliferum. Sarsia 81:339–344. https://doi.org/10.1080/00364827.1996.10413631
Tillmann U (2003) Kill and eat your predator: a winning strategy of the planktonic flagellate Prymnesium parvum. Aq Microb Ecol 32:73–84. https://doi.org/10.3354/ame032073
Jones AC, Liao TSV, Najar FZ, Roe BA, Hambright KD, Caron DA (2013) Seasonality and disturbance: annual pattern and response of the bacterial and microbial eukaryotic assemblages in a freshwater ecosystem. Environ. Microbiol. 15:2557–2572. https://doi.org/10.1111/1462-2920.12151
Amaral-Zettler LA, McCliment EA, Ducklow HW, Huse SM (2009) A method for studying protistan diversity using massively parallel sequencing of V9 hypervariable regions of small-subunit ribosomal RNA genes. PLoS One 4:e6372
Sogin ML, Morrison HG, Huber JA, Welch DM, Huse SM, Neal PR, Arrieta JM, Herndl GJ (2006) Microbial diversity in the deep sea and the underexplored “rare biosphere”. Proc. Natl. Acad. Sci. U. S. A. 103:12115–12120
Huse S, Huber J, Morrison H, Sogin M, Welch D (2007) Accuracy and quality of massively parallel DNA pyrosequencing. Genome Biol. 8:R143
Huse SM, Welch DM, Morrison HG, Sogin ML (2010) Ironing out the wrinkles in the rare biosphere through improved OTU clustering. Environ. Microbiol. 12:1889–1898
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF (2009) Introducing Mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75:7537–7541. https://doi.org/10.1128/aem.01541-09
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25:3389–3402
Pruesse E, Quast C, Knittel K, Fuchs B, Ludwig W, Peplies J, Glöckner FO (2007) SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res. 35:7188–7196
Clarke KR (1993) Non-parametric multivariate analyses of changes in community structure. Australian J Ecol 18: 117–143
Clarke KR, Warwick RM (2001) Change in marine communities: an approach to statistical analysis and interpretation, 2nd, Plymouth, UK
Chow C-ET, Kim DY, Sachdeva R, Caron DA, Fuhrman JA (2014) Top-down controls on bacterial community structure: microbial network analysis of bacteria, T4-like viruses and protists. ISME J 8:816–829
Ruan Q, Dutta D, Schwalbach MS, Steele JA, Fuhrman JA, Sun F (2006) Local similarity analysis reveals unique associations among marine bacterioplankton species and environmental factors. Bioinformatics 22:2532–2538. https://doi.org/10.1093/bioinformatics/btl417
Storey JD (2002) A direct approach to false discovery rates. J Royal Stat Soc: Series B (Stat Methodol) 64:479–498. https://doi.org/10.1111/1467-9868.00346
Xia L, Steele J, Cram J, Cardon Z, Simmons S, Vallino J, Fuhrman J, Sun F (2011) Extended local similarity analysis (eLSA) of microbial community and other time series data with replicates. BMC Systems Biol 5:S15
Cline MS, Smoot M, Cerami E, Kuchinsky A, Landys N, Workman C, Christmas R, Avila-Campilo I, Creech M, Gross B, Hanspers K, Isserlin R, Kelley R, Killcoyne S, Lotia S, Maere S, Morris J, Ono K, Pavlovic V, Pico AR, Vailaya A, Wang P-L, Adler A, Conklin BR, Hood L, Kuiper M, Sander C, Schmulevich I, Schwikowski B, Warner GJ, Ideker T, Bader GD (2007) Integration of biological networks and gene expression data using Cytoscape. Nat. Protoc. 2:2366–2382
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13:2498–2504. https://doi.org/10.1101/gr.1239303
Smoot ME, Ono K, Ruscheinski J, Wang P-L, Ideker T (2011) Cytoscape 2.8: new features for data integration and network visualization. Bioinformatics 27:431–432. https://doi.org/10.1093/bioinformatics/btq675
Assenov Y, Ramírez F, Schelhorn S-E, Lengauer T, Albrecht M (2008) Computing topological parameters of biological networks. Bioinformatics 24:282–284. https://doi.org/10.1093/bioinformatics/btm554
Erdös P, Réyni A (1960) On the evolution of random graphs. Inst Math Hungarian Acad Sci 5:17–61
Bader G, Hogue C (2003) An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinform 4:2
Jost L (2006) Entropy and diversity. Oikos 113:363–375
Friedman J, Alm EJ (2012) Inferring correlation networks from genomic survey data. PLoS Comp Biol 8(9):e1002687
Barabasi A-L, Oltvai ZN (2004) Network biology: understanding the cell’s functional organization. Nature Rev Gen 5:101–113
Lima-Mendez G, van Helden J (2009) The powerful law of the power law and other myths in network biology. Molec BioSys 5:1482–1493
de Vargas C, Audic S, Henry N, Decelle J, Mahé F, Logares R, Lara E, Berney C, Le Bescot N, Probert I, Carmichael M, Poulain J, Romac S, Colin S, Aury J-M, Bittner L, Chaffron S, Dunthorn M, Engelen S, Flegontova O, Guidi L, Horák A, Jaillon O, Lima-Mendez G, Lukeš J, Malviya S, Morard R, Mulot M, Scalco E, Siano R, Vincent F, Zingone A, Dimier C, Picheral M, Searson S, Kandels-Lewis S, Coordinators TO, Acinas SG, Bork P, Bowler C, Gorsky G, Grimsley N, Hingamp P, Iudicone D, Not F, Ogata H, Pesant S, Raes J, Sieracki ME, Speich S, Stemmann L, Sunagawa S, Weissenbach J, Wincker P, Karsenti E (2015) Eukaryotic plankton diversity in the sunlit ocean. Science 348:1261605. https://doi.org/10.1126/science.1261605
Watts DJ, Strogatz SH (1998) Collective dynamics of “small-world” networks. Nature 393:440–442
Gobler CJ, Sunda WG (2012) Ecosystem disruptive algal blooms of the brown tide species, Aureococcus anophagefferens and Aureoumbra lagunensis. Harmful Algae 14:36–45
Igarashi T, Satake M, Yasumoto T (1996) Prymnesin-2: a potent ichthyotoxic and hemolytic glycoside isolated from the red tide alga Prymnesium parvum. J. Am. Chem. Soc. 118:479–480. https://doi.org/10.1021/ja9534112
Hambright KD, Beyer JE, Easton JD, Zamor RM, Easton AC, Hallidayschult TC (2015) The niche of an invasive marine microbe in a subtropical freshwater impoundment. ISME J 9:256–264. https://doi.org/10.1038/ismej.2014.103
Roelke DL, Barkoh A, Brooks BW, Grover JP, Hambright KD, LaClaire JW, Moeller PDR, Patino R (2015) A chronicle of a killer alga in the west: ecology, assessment, and management of Prymnesium parvum blooms. Hydrobiologia 764:29–50. https://doi.org/10.1007/s10750-015-2273-6
Acosta F, Zamor R, Najar F, Roe B, Hambright KD (2015) Dynamics of an experimental microbial invasion. Proc. Natl. Acad. Sci. U. S. A. 112:11594–11599
Michaloudi E, Moustaka-Gouni M, Gkelis S, Pantelidakis K (2008) Plankton community structure during an ecosystem disruptive algal bloom of Prymnesium parvum. J. Plankton Res. 31:301–309
Acknowledgements
The authors would like to thank James D. Easton, Anne C. Easton, and Richard Zamor for field measurements, sample collection and microscopical counts, and Bruce Roe and Fares Z. Najar for DNA sequencing. All sequences are located in the NCBI short read archive under project BioProject PRJNA195945.
Funding
Funding was provided by a grant from the Oklahoma Department of Wildlife Conservation through the Sport Fish Restoration Program (grant F-61-R) to KDH. Oklahoma Mesonet data are provided courtesy of the Oklahoma Mesonet, which is jointly operated by Oklahoma State University and the University of Oklahoma. Continued funding for maintenance of the observing network is provided by the taxpayers of Oklahoma.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
ESM 1
(PDF 82 kb)
ESM 2
(PDF 279 kb)
ESM 3
(PDF 41 kb)
ESM 4
(PDF 102 kb)
ESM 5
(PDF 67 kb)
Fig. S1
Graphs depicting the annual trajectories of P. parvum cell counts (a), and Prymnesium-affiliated OTUs (b and c) in Lebanon Pool (A and B) and Wilson Creek (a and c). Panel a shows the abundances of P. parvum cell counts in Lebanon Pool (black symbols and lines; left Y axis) and Wilson Creek (grey symbols and lines; right Y axis). Note different scales. Panels b and c show the relative abundances of 18S OTU1 (solid symbols), plus 16S OTU1 and 16S OTU12 (open symbols) in Lebanon Pool (b) and Wilson Creek (c). Note the different scales in (b) and (c). (JPEG 33 kb)
Fig. S2
Histograms of all the permuted p-values associated with all Spearman correlations from a) Lebanon Pool and b) Wilson Creek. (JPEG 23 kb)
Fig. S3
The frequency, on a log scale, of a microbial OTUs relative abundance plotted against its number of significant Spearman correlations with other OTUs or environmental variables for the data from Lebanon Pool (a) and Wilson Creek (b). Lines indicate the 95% confidence intervals for the number of correlations per OTU, and its relative abundances. The boxes highlight OTUs of interest: 1) the open boxes outlined in gray, in the upper right contain OTUs with large (outside the 95% CI) relative abundances and numbers of significant correlations; 2) the shaded boxes on the upper left contain OTUs with large (outside the 95% CI) relative abundances and small (outside the 95% CI) numbers of significant correlations; and 3) the shaded box on the lower right contains OTUs with small (outside the 95% CI) relative abundances and large (outside the 95% CI) numbers of significant correlations. The average number of correlations per OTU in Lebanon Pool was 43 (Std. Dev = 31) and the average in Wilson Creek was 31 (Std. Dev = 24). There were OTUs with high average relative abundances and a large number of correlations (A and B, open gray boxes) for example: in Lebanon Pool a Prymnesium plastid (16S_1 = 6.7% average abundance and 10 occurrences), a diatom plastid (18S_10 = 3% average abundance and 12 occurrences), and a SAR11 (OTU_4 = 1.8% average abundance and 12 occurrences), each had over 87 correlations. In addition, we observed nodes that fell outside the 95% confidence intervals for number of correlations and average relative abundances (upper left shaded boxes). Two fungal OTUs (18S_5 and 18S_168), one in Lebanon Pool and both in Wilson Creek were highly abundant yet had fewer than 8 correlations each (upper left shaded boxes). In contrast, in Wilson Creek (lower right shaded box), a chlorophyte (18S_190), a ciliate (18S_223), and a fungal (18S_288) OTU each had low relative abundances but a high (> 70) number of significant correlations. (JPEG 34 kb)
Fig. S4
Log distributions representing the number of significant spearman correlations per microbial OTU or environmental variable within the networks from: a) Lebanon Pool, b) Wilson Creek, and c) shared at both locations. Closed symbols represent the distribution from the experimental microbial association networks, and open symbols represent distributions constructed from Erdös-Réyni model networks of the same size as the experimental networks. The upper inset graphs in each panel show Poisson distributions fit to the Erdös-Réyni model data with r2s of: a) Lebanon Pool = 0.80, b) Wilson Creek = 0.87, and c) shared = 0.87. The distribution for the shared microbial association network (c, lower inset) had a moderate fit r2 of 0.6 to a power curve. (JPEG 29 kb)
Fig. S5
Spearman correlations between the nodes within the network diagrams revealed seasonal abundance patterns among the microbial taxa shared at both locations. The networks were visualized with the unweighted force-directed layout (nodes in the network were positioned based on the number of Spearman correlations). OTUs with 75% or more of their relative abundances contained in the six month period of November-April (c) or May-October (d) are highlighted in yellow. Connections drawn from positive Spearman correlations are black solid lines, and those from negative correlations are gray dotted lines. All correlations (510 [>0.7 or <-0.7 and p-values ≤0.01]) are displayed in panels a, c and d. Only positive correlations (353) are displayed in panel b. Bacteria are red circles, eukaryotes are blue or purple (metazoa) diamonds, environmental parameters are orange squares, and chloroplasts are green circles. (JPEG 49 kb)
Fig. S6
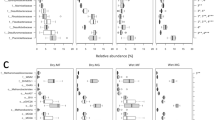
Network representations of selected individual positive Spearman correlations between a plastid’s OTUs and its likely photosynthetic eukaryotic host (i.e. 18S OTU or P. parvum cell count) are shown for the data from Lebanon Pool (a) and Wilson Creek (b). Connections represent positive Spearman correlations (>0.7 and p-values ≤0.01) and the exact values are written on the lines. Single-celled eukaryotes are blue diamonds, environmental parameters are orange squares, and chloroplasts are green circles. The size of the symbol reflects the average relative sequence abundance. The number on the symbols refers to the OTU identifier numbers. The following identification codes were used for the OTUs with good taxonomic resolution: Hap (haptophyte), Chr (chrysophyte), Chl (chlorophyte), Cry (cryptophyte), Dia (diatom), and Eug (euglenid), Dict (dictyophyte), UC (unclassified), Ppar (P. parvum cell counts). Refer to Tables S1 and S2 for a complete list of the OTUs and their identifications. (JPEG 89 kb)
Rights and permissions
About this article
Cite this article
Jones, A.C., Hambright, K.D. & Caron, D.A. Ecological Patterns Among Bacteria and Microbial Eukaryotes Derived from Network Analyses in a Low-Salinity Lake. Microb Ecol 75, 917–929 (2018). https://doi.org/10.1007/s00248-017-1087-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-017-1087-7