Abstract
Isozymes, homologous enzymes coded by separate loci within a genome, present interesting systems for examining molecular and functional divergence through natural selection. Isozyme pairs for a number of metabolic enzymes, including Triosephosphate isomerase (Tpi), Malate dehydrogenase (Mdh), Phosphoglucose isomerase (Pgi), and Guanylate kinase (Guk), appear to all result from a single, large duplication event early in teleost evolution. These small gene families include two forms, a generally expressed form with no apparent charge and a neurally expressed form with a pronounced negative charge although the canalization of expression of the second form varies across families. Using ancestral sequence reconstructions and standard comparisons of rates of nonsynonymous and synonymous change, combined with the examination of the specific amino acid changes observed and predicted we examined the evolution of the Tpi and Guk families using all available vertebrate sequences and all four families using a smaller, common, dataset. We find that post-duplication, the neural Tpi and Guk isozymes evolved through similar periods of positive selection as evidenced by elevated rates of nonsynonymous change and accumulation of negative amino acids. Over the same evolutionary period our analysis suggests that Mdh and Pgi isozymes appear to have evolved under a less divergent pattern of selection. These distinct results likely reflect functional differences between the isozymes, possibly a result of differences in expression patterns.





Similar content being viewed by others
References
Agarwal KC, Miech RP, Parks RE Jr (1978) Guanylate kinases from human erythrocytes, hog brain, and rat liver. Methods Enzymol 51:483–490
Altschul S, Gish W, Miller W, Myers E, Lipman D (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410
Amores A, Force A, Yan Y, Joly L, Amemiya C, Fritz A, Ho RK, Langeland J, Prince V, Wang Y, Westerfield M, Ekker M, Postlethwait JH (1998) Zebrafish hox clusters and vertebrate genome evolution. Science 282(27):1711–1714
Anisimova M, Yang Z (2007) Multiple hypothesis testing to detect lineages under positive selection that affects only a few sites. Mol Biol Evol 24(5):1219–1228
Beisswanger S, Stephan W (2008) Evidence that strong positive selection drives neofunctionalization in the tandemly duplicated polyhomeotic genes in Drosophila. Proc Natl Acad Sci USA 105(14):5447–5452
Bollback JP (2002) Bayesian model adequacy and choice in phylogenetics. Mol Biol Evol 19(7):1171–1180
Champion MJ, Whitt GS (1976) Differential gene expression in multilocus isozyme systems of the developing green sunfish. J Exp Zool 196(3):263–281
Charlesworth B, Morgan MT, Charlesworth D (1993) The effect of deleterious mutations on neutral molecular variation. Genetics 134:1289–1303
Chiu C-h, Dewar K, Wagner GP, Takahashi K, Ruddle F, Ledje C, Bartsch P, Scemama J-L, Stellwag E, Fried C, Prohaska SJ, Stadler PF, Amemiya CT (2004) Bichir HoxA cluster sequence reveals surprising trends in ray-finned fish genomic evolution. Genome Res 14(1):11–17
Daly LE, Bourke GJ (2008) Interpretation and uses of medical statistics, 5th edn. Blackwell Science, Oxford
Douard V, Brunet F, Boussau B, Ahrens-Fath I, Vlaeminck-Guillem V, Haendler B, Laudet V, Guiguen Y (2008) The fate of the duplicated androgen receptor in fishes: a late neofunctionalization event? BMC Evol Biol 8(1):336
Dykhuizen D, Hartl DL (1980) Selective neutrality of 6PGD allozymes in E. coli and the effects of genetic background. Genetics 96:801–817
Elkin BS, Shaik MA, Morrison B (2010) Fixed negative charge and the donnan effect: a description of the driving forces associated with brain tissue swelling and oedema. Philos Trans R Soc A 368(1912):585–603
Felsenstein J (1989) PHYLIP—phylogeny inference package. Cladistics 5 Version 3.2:164–166
Felsenstein J (1978) Cases in which parsimony or compatibility methods will be positively misleading. Syst Zool 27(4):401–410
Fisher SE, Whitt GS (1978) Evolution of isozyme loci and their differential tissue expression creatine kinase as a model system. J Mol Evol 12(1):25–55
Fisher SE, Shaklee JB, Ferris SD, Whitt GS (1980) Evolution of five multilocus isozyme systems in the chordates. Genetica 52–53(1):73–85
Gaida IH (1995) Evolutionary aspects of gene expression in the Pacific Angel Shark, Squatina California (Squatiniformes: Squatinidae). Copeia 3:532–554
Goodman M, Moore GW, Matsuda G (1975) Darwinian evolution in the genealogy of haemoglobin. Nature 253(5493):603–608
He X, Zhang J (2005) Rapid subfunctionalization accompanied by prolonged and substantial neofunctionalization in duplicate gene evolution. Genetics 169(2):1157–1164
Huelsenbeck JP (1995) The robustness of two phylogenetic methods: four-taxon simulations reveal a slight superiority of maximum likelihood over neighbor joining. Mol Biol Evol 12(5):843–849
Huelsenbeck JP, Hillis DM (1993) Success of phylogenetic methods in the four-taxon case. Syst Biol 42(3):247–264
Hughes AL (1994) The evolution of functionally novel proteins after gene duplication. Proc R Soc B 256(1346):119–124
Hughes AL, Friedman R (2008) Codon-based tests of positive selection, branch lengths, and the evolution of mammalian immune system genes. Immunogenetics 60(9):495–506
Jaillon O, Aury J, Brunet F, Petit J, Stange-Thomann N, Mauceli E, Bouneau L, Fischer C, Ozouf-Costaz C, Bernot A, Nicaud S, Jaffe D, Fisher S, Lutfalla G, Dossat C, Segurens B, Dasilva C, Salanoubat M, Levy M, Boudet N, Castellano S, Anthouard V, Jubin C, Castelli V, Katinka M, Vacherie B, Biémont C, Skalli Z, Cattolico L, Poulain J, De Berardinis V, Cruaud C, Duprat S, Brottier P, Coutanceau J, Gouzy J, Parra G, Lardier G, Chapple C, McKernan KJ, McEwan P, Bosak S, Kellis M, Volff J, Guigó R, Zody MC, Mesirov J, Lindblad-Toh K, Birren B, Nusbaum C, Kahn D, Robinson-Rechavi M, Laudet V, Schachter V, Quétier F, Saurin W, Scarpelli C, Wincker P, Lander ES, Weissenbach J, Roest Crollius H (2004) Genome duplication in the teleost fish Tetraodon nigroviridis reveals the early vertebrate proto-karyotype. Nature 431(7011):946–957
Kellis M, Birren BW, Lander ES (2004) Proof and evolutionary analysis of ancient genome duplication in the yeast Saccharomyces cerevisiae. Nature 428(6983):617–624
Kettler MK, Whitt GS (1986) An apparent progressive and recurrent evolutionary restriction in tissue expression of a gene, the lactate dehydrogenase-c gene, within a family of bony fish (Salmoniformes: Umbridae). J Mol Evol 23(2):95–107
Kimura M (1983) The neutral theory of molecular evolution. Cambridge University Press, Cambridge
King JL, Jukes TH (1969) Non-Darwinian evolution. Science 164:788–798
Kosiol C, Vinař T, Da Fonseca RR, Hubisz MJ, Bustamante CD, Nielsen R, Siepel A (2008) Patterns of positive selection in six mammalian genomes. PLoS Genet 4(8):E1000144–E1000416
Li WH, Gojobori T (1983) Rapid evolution of goat and sheep globin genes following gene duplication. Mol Biol Evol 1(1):94–108
Lynch VJ (2007). Inventing an arsenal: adaptive evolution and neofunctionalization of snake venom phospholipase A2 genes. BMC Evol Biol 7(2). doi:10.1186/1471-2148-7-2
Lynch M, Force A (2000) The probability of duplicate gene preservation by subfunctionalization. Genetics 154(1):459–473
Margolis RK, Margolis RU (1993) Nervous tissue proteoglycans. Experientia 49(5):429–446
Markert CL, Shaklee JB, Whitt GS (1975) Evolution of a gene. Science 189:102–114
Merritt TJS, Quattro JM (2001) Evidence for a period of directional selection following gene duplication in a neurally expressed locus of triosephosphate isomerase. Genetics 159:689–697
Merritt TJS, Quattro JM (2002) Negative charge correlates with neural expression in vertebrate aldolase isozymes. J Mol Evol 55(6):674–683
Merritt TJS, Quattro JM (2003) Evolution of the vertebrate cytosolic malate dehydrogenase gene family: duplication and divergence in actinopterygian fish. J Mol Evol 56(3):265–276
Messier W, Stewart C (1997) Episodic adaptive evolution of primate lysozymes. Nature 385(6612):151–154
Meyer A, Van De Peer Y (2005) From 2R to 3R: evidence for a fish-specific genome duplication (FSGD). BioEssays 27(9):937–945
Moore BW (1973) Brain specific proteins. In: Proteins of the nervous system. Raven, New York, pp 1–12
Moore BW, McGregor D (1965) Chromatographic and electrophoretic fractionation of soluble proteins of brain and liver. J Biol Chem 240:1647–1653
Morizot DC, Slaugenhaupt SA, Kallman KD, Chakravarti A (1991) Genetic linkage map of fishes of the genus Xiphophorus (Teleostei: Poeciliidae). Genetics 127(2):399–410
Nei M, Kumar S (2000) Molecular evolution and phylogenetics. Oxford University Press, Oxford
Novak U, Kaye AH (2000) Extracellular matrix and the brain: components and function. J Clin Neurosci 7(4):280–290
Ohno S (1970) Evolution by gene duplication. Springer, Berlin
Ohno S, Wolf U, Atkin NB (1968) Evolution from fish to mammals by gene duplication. Hereditas 59(1):169–187
Ohta T (1991) Multigene families and the evolution of complexity. J Mol Evol 33(1):34–41
Pebusque MJ, Coulier F, Birnbaum D, Pontarotti P (1998) Ancient large-scale genome duplications: phylogenetic and linkage analyses shed light on chordate genome evolution. Mol Biol Evol 15(9):1145–1159
Philippe H (2000) Opinion: long branch attraction and protist phylogeny. Protist 151:307–316
Pommie C, Levadoux S, Sabatier R, Lefranc G, Lefranc MP (2004) IMGT standardized criteria for statistical analysis of immunoglobulin V-REGION amino acid properties. J Mol Recognit 17:17–32
Pontier PJ, Hart NH (1981) Developmental expression of glucose and triose phosphate isomerase genes in teleost fishes (Brachydanio). J Exp Zool 217(1):53–71
Postlethwait JH, Yan Y, Gates MA, Horne S, Amores A, Brownlie A, Donovan A, Egan ES, Force A, Gong Z, Goutel C, Fritz A, Kelsh R, Knapik E, Liao E, Paw B, Ransom D, Singer A, Thomson M, Abduljabbar TS, Yelick P, Beier D, Joly J-S, Larhammar D, Rosa F, Westerfield M, Zon LI, Johnson SL, Talbot WS (1998) Vertebrate genome evolution and the Zebrafish gene map. Nat Genet 18(4):345–349
Quattro JM, Woods HA, Powers DA (1993) Sequence analysis of teleost retina-specific lactate dehydrogenase C: evolutionary implications for the vertebrate lactate dehydrogenase gene family. Proc Natl Acad Sci USA 90(1):242–246
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19(12):1572–1574
Sato Y, Nishida M (2007) Post-duplication charge evolution of phosphoglucose isomerases in teleost fishes through weak selection on many amino acid sites. BMC Evol Biol 7(1):204
Shaklee JB, Kepes KL, Whitt GS (1973) Specialized lactate dehydrogenase isozymes: the molecular and genetic basis for the unique eye and liver LDHS of teleost fishes. J Exp Zool 185(2):217–240
Soltis PS, Soltis DE (2000) The role of genetic and genomic attributes in the success of polyploids. Proc Natl Acad Sci USA 97(13):7051–7057
Steinke D, Hoegg S, Brinkmann H, Meyer A (2006) Three rounds (1R/2R/3R) of genome duplications and the evolution of the glycolytic pathway in vertebrates. BMC Biol 4(16):16
Stothard P (2000) The Sequence Manipulation Suite: JavaScript programs for analyzing and formatting protein and DNA sequences. Biotechniques 28:1102–1104
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24(8):1596–1599
Taylor JS, De Peer Y, Braasch I, Meyer A (2001) Comparative genomics provides evidence for an ancient genome duplication event in fish. Philos Trans R Soc B 356(1414):1661–1679
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22(22):4673–4680
Vandepoele K, De Vos W, Taylor JS, Meyer A, Van De Peer Y (2004) Major events in the genome evolution of vertebrates: paranome age and size differ considerably between ray-finned fishes and land vertebrates. Proc Natl Acad Sci USA 101(6):1638–1643
Volff J-N (2005) Genome evolution and biodiversity in teleost fish. Heredity 94(3):280–294
Wendel JF (2000) Genome evolution in polyploids. Plant Mol Biol 42(1):225–249
Whitt GS (1970) Developmental genetics of the lactate dehydrogenase isozymes of fish. J Exp Zool 175(1):1–35
Whitt GS (1983) Isozymes as probes and participants in developmental and evolutionary genetics. Isozymes Curr Top Biol Med Res 10:1–40
Yang Z (1998) Likelihood ratio tests for detecting positive selection and application to primate lysozyme evolution. Mol Biol Evol 15(5):568–573
Yang Z (2002) Inference of selection from multiple species alignments. Curr Opin Genet Dev 12(6):688–694
Yang Z (2007) PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24(8):1586–1591
Yang Z, Bielawski JP (2000) Statistical methods for detecting molecular adaptation. Trends Ecol Evol 15(12):496–503
Yang Z, Goldman N, Friday A (1994) Comparison of models for nucleotide substitution used in maximum-likelihood phylogenetic estimation. Mol Biol Evol 11(2):316–324
Yang Z, Kumar S, Nei M (1995) A new method of inference of ancestral nucleotide and amino acid sequences. Genetics 141:1641–1650
Yokoyama S, Tada T, Zhang H, Britt L (2008) From the cover: elucidation of phenotypic adaptations: molecular analyses of dim-light vision proteins in vertebrates. Proc Natl Acad Sci USA 105(36):13480–13485
Zhang J, Nei M (1997) Accuracies of ancestral amino acid sequences inferred by the parsimony, likelihood, and distance methods. J Mol Evol 44:139–146
Zhang J, Kumar S, Nei M (1997) Small-sample tests of episodic adaptive evolution: a case study of primate lysozymes. Mol Biol Evol 14(12):1335–1338
Zhang J, Rosenberg HF, Nei M (1998) Positive Darwinian selection after gene duplication in primate ribonuclease genes. Proc Natl Acad Sci USA 95:3708–3713
Acknowledgments
We thank Teresa Rzezniczak, David Bing, Jose Knee, Brendan McConkey, Katharine Coykendall, and Efe Sezgin for constructive comments on this manuscript, as well as Jennifer Fenske and Stefan Siemann for phylogenetic and protein structure assistance. This study was supported by the Natural Sciences and Engineering Research Council of Canada (NSERC) Discovery (3414-07) and a Canada Research Chair (950-215763) to TJSM.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Online Resource Fig. 1
Phylogenetic tree produced from the large Tpi nucleotide sequence dataset by NJ, Bayesian, and ML methods. Above each branch, proportions of 1,000 bootstrap replicates for NJ, Bayesian, and ML methods are shown. Asterisk represent nodes without bootstrap support due to separate topology being inferred by given phylogenetic method (see “Results” section)
Online Resource Fig. 2
Phylogenetic tree produced from the large Guk nucleotide sequence dataset by NJ, Bayesian, and ML methods. Above each branch, proportions of 1,000 bootstrap replicates for NJ, Bayesian, and ML methods are shown. Asterisk represent nodes without bootstrap support due to separate topology being inferred by the given phylogenetic method. The given topology is consistent with what is known from the related gene families in the teleost family, such as Tpi, Mdh, and Pgi
Online Resource Fig. 3
Phylogenetic trees of comparison datasets by NJ, Bayesian, and ML methods with 1,000 bootstrap replicate support respectively along each branch for Tpi (a), Guk (b), Mdh (c), and Pgi (d). Both Guk and Tpi comparison dataset produce identical topologies by all three phylogenetic methods, while Mdh and Pgi produced the shown topologies for two of the three phylogenetic methods. Asterisk represents nodes that were unresolved by the given phylogenetic methods
Online Resource Fig. 4
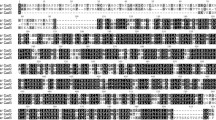
(Tpi) Inferred amino acid sequences for the duplication node (AncestralS), node leading to the neural (AncestralN), and general (AncestralG) isozyme branches of the common Tpi tree. Highlighted amino acids were found to reside on the outside of the protein by solvent accessibility of >25 %. Asterisks indicate amino acid substitutions toward negative amino acids, and isoelectric points are shown as the 3′ end of the proteins
Online Resource Fig. 5
(Mdh) Inferred amino acid sequences for the duplication node (AncestralS), node leading to the neural (AncestralN), and general (AncestralG) isozyme branches of the common Mdh tree. Highlighted amino acids were found to reside on the outside of the protein by solvent accessibility of >25 %. Asterisks indicate amino acid substitutions toward negative amino acids, and isoelectric points are shown as the 3′ end of the proteins
Rights and permissions
About this article
Cite this article
Auld, R.R., Quattro, J.M. & Merritt, T.J.S. Molecular Evolution of Teleost Neural Isozymes. J Mol Evol 75, 198–213 (2012). https://doi.org/10.1007/s00239-012-9532-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-012-9532-1