Abstract
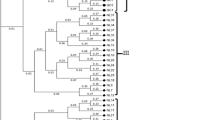
Segregation and linkage relationships were analyzed between 28 isoenzyme loci in ten natural stands representing much of the natural range of Pinus echinata Mill. (shortleaf pine). A total of 203 possible two-locus combinations were tested. Three linkage groups were revealed in this study at a linkLOD of 4.0. The first linkage group (A) consisted of Pgi and Adh-1; Gdh, Idh, Skdh-2, G6pd-2 and Aco were mapped to the second linkage group (B); the third group (C) had 2 loci: Mdh-2 and Mdh-3. A moderate linkage between Mnr-2 and Dia-2 and weak linkages between Mnr-1 and Dia-1, and Got-2 and 6pgd-2 were also detected. The significance of these results in shortleaf pine is discussed and compared to linkage maps previously reported in other conifers, including pines.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Received: 7 April 1997 / Accepted: 25 April 1997
Rights and permissions
About this article
Cite this article
Raja, R., Tauer, C., Wittwer, R. et al. Segregation and linkage relationships of isoenzymes in shortleaf pine (Pinus echinata Mill.). Theor Appl Genet 95, 1252–1257 (1997). https://doi.org/10.1007/s001220050689
Issue Date:
DOI: https://doi.org/10.1007/s001220050689