Abstract
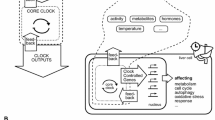
Circadian timekeeping is a ubiquitous mechanism that enables organisms to maintain temporal coordination between internal biological processes and time of the local environment. The molecular basis of circadian rhythms lies in a set of transcription–translation feedback loops (TTFLs) that drives the rhythmic transcription of core clock genes, whose level and phase of expression serve as the marker of circadian time. However, it has become increasingly evident that additional regulatory mechanisms impinge upon the TTFLs to govern the properties and behavior of the circadian clock. Such mechanisms include changes in chromatin architecture, interactions with other transcription factor networks, post-transcriptional control by RNA modifications, alternative splicing and microRNAs, and post-translational regulation of subcellular trafficking and protein degradation. In this review, we will summarize the current knowledge of circadian clock regulation—from transcriptional to post-translational—drawing from literature pertaining to the Drosophila and murine circadian systems.




Similar content being viewed by others
References
Abrahamson EE, Moore RY (2001) Suprachiasmatic nucleus in the mouse: retinal innervation, intrinsic organization and efferent projections. Brain Res 916:172–191. doi:10.1016/S0006-8993(01)02890-6
Maywood ES, Chesham JE, O’Brien JA, Hastings MH (2011) A diversity of paracrine signals sustains molecular circadian cycling in suprachiasmatic nucleus circuits. Proc Natl Acad Sci USA 108:14306–14311. doi:10.1073/pnas.1101767108
Nitabach MN, Taghert PH (2008) Organization of the Drosophila circadian control circuit. Curr Biol 18:R84–R93. doi:10.1016/j.cub.2007.11.061
Renn SC, Park JH, Rosbash M, Hall JC, Taghert PH (1999) A pdf neuropeptide gene mutation and ablation of PDF neurons each cause severe abnormalities of behavioral circadian rhythms in Drosophila. Cell 99:791–802. doi:10.1016/S0092-8674(00)81676-1
Lin Y, Stormo GD, Taghert PH (2004) The neuropeptide pigment-dispersing factor coordinates pacemaker interactions in the Drosophila circadian system. J Neurosci 24:7951–7957. doi:10.1523/JNEUROSCI.2370-04.2004
Hall JC (2003) Genetics and molecular biology of rhythms in Drosophila and other insects. Adv Genet 48:1–280. doi:10.1016/S0065-2660(03)48000-0
Ashmore LJ, Sehgal A (2003) A fly’s eye view of circadian entrainment. J Biol Rhythms 18:206–216. doi:10.1177/0748730403253385
Helfrich-Förster C, Winter C, Hofbauer A, Hall JC, Stanewsky R (2001) The circadian clock of fruit flies is blind after elimination of all known photoreceptors. Neuron 30:249–261. doi:10.1016/S0896-6273(01)00277-X
Emery P, So WV, Kaneko M, Hall JC, Rosbash M (1998) CRY, a Drosophila clock and light-regulated cryptochrome, is a major contributor to circadian rhythm resetting and photosensitivity. Cell 95:669–679. doi:10.1016/S0092-8674(00)81637-2
Schibler U, Gotic I, Saini C, Gos P, Curie T, Emmenegger Y, Sinturel F, Gosselin P, Gerber A, Fleury-Olela F, Rando G, Demarque M, Franken P (2015) Clock-talk: interactions between central and peripheral circadian oscillators in mammals. Cold Spring Harb Sym 80:223–232. doi:10.1101/sqb.2015.80.027490
Poletini MO, Moraes MN, Ramos BC, Jerônimo R, Castrucci AM (2015) TRP channels: a missing bond in the entrainment mechanism of peripheral clocks throughout evolution. Temperature 2:522–534. doi:10.1080/23328940.2015.1115803
Tahara Y, Aoyama S, Shibata S (2016) The mammalian circadian clock and its entrainment by stress and exercise. J Physiol Sci. doi:10.1007/s12576-016-0450-7
Mehta N, Cheng H-YM (2013) Micro-managing the circadian clock: the role of microRNAs in biological timekeeping. J Mol Biol 425:3609–3624. doi:10.1016/j.jmb.2012.10.022
Lowrey PL, Takahashi JS (2011) Genetics of circadian rhythms in mammalian model organisms. Adv Genet 74:175–230. doi:10.1016/B978-0-12-387690-4.00006-4
Landgraf D, Wang LL, Diemer T, Welsh DK (2016) NPAS2 compensates for loss of CLOCK in peripheral circadian oscillators. PLoS Genet 12:e1005882. doi:10.1371/journal.pgen.1005882
Koike N, Yoo SH, Huang HC, Kumar V, Lee C, Kim TK, Takahashi JS (2012) Transcriptional architecture and chromatin landscape of the core circadian clock in mammals. Science 338:349–354. doi:10.1126/science.1226339
Ye R, Selby CP, Chiou YY, Ozkan-Dagliyan I, Gaddameedhi S, Sancar A (2014) Dual modes of CLOCK:BMAL1 inhibition mediated by Cryptochrome and Period proteins in the mammalian circadian clock. Gene Dev 28:1989–1998. doi:10.1101/gad.249417.114
Hardin PE (2011) Molecular genetic analysis of circadian timekeeping in Drosophila. Adv Genet 74:141–173. doi:10.1016/B978-0-12-387690-4.00005-2
Kadener S, Stoleru D, McDonald M, Nawathean P, Rosbash M (2007) Clockwork Orange is a transcriptional repressor and a new Drosophila circadian pacemaker component. Gene Dev 21:1675–1686. doi:10.1101/gad.1552607
Matsumoto A, Ukai-Tadenuma M, Yamada RG, Houl J, Uno KD, Kasukawa T, Dauwalder B, Itoh TQ, Takahashi K, Ueda R, Hardin PE, Tanimura T, Ueda HR (2007) A functional genomics strategy reveals clockwork orange as a transcriptional regulator in the Drosophila circadian clock. Gene Dev 21:1687–1700. doi:10.1101/gad.1552207
Roenneberg T, Merrow M (2016) The circadian clock and human health. Curr Biol 26:R432–R443. doi:10.1016/j.cub.2016.04.011
García-González E, Escamilla-Del-Arenal M, Arzate-Mejía R, Recillas-Targa F (2016) Chromatin remodeling effects on enhancer activity. Cell Mol Life Sci 73:2897–2910. doi:10.1007/s00018-016-2184-3
Groth A, Rocha W, Verreault A, Almouzni G (2007) Chromatin challenges during DNA replication and repair. Cell 128:721–733. doi:10.1016/j.cell.2007.01.030
Bannister AJ, Kouzarides T (2011) Regulation of chromatin by histone modifications. Cell Res 21:381–395. doi:10.1038/cr.2011.22
Shahbazian MD, Grunstein M (2007) Functions of site-specific histone acetylation and deacetylation. Annu Rev Biochem 76:75–100. doi:10.1146/annurev.biochem.76.052705.162114
Schmitt A, Gutierrez GJ, Lénárt P, Ellenberg J, Nebreda AR (2002) Histone H3 phosphorylation during Xenopus oocyte maturation: regulation by the MAP kinase/p90Rsk pathway and uncoupling from DNA condensation. FEBS Lett 518:23–28. doi:10.1016/S0014-5793(02)02630-3
Dong Z, Bode AM (2006) The role of histone H3 phosphorylation (Ser10 and Ser28) in cell growth and cell transformation. Mol Carcinog 45:416–421. doi:10.1002/mc.20220
Pal S, Sif S (2007) Interplay between chromatin remodelers and protein arginine methyltransferases. J Cell Physiol 213:306–315. doi:10.1002/jcp.21180
Kouzarides T (2007) Chromatin modifications and their function. Cell 128:693–705. doi:10.1016/j.cell.2007.02.005
Crosio C, Cermakian N, Allis CD, Sassone-Corsi P (2000) Light induces chromatin modification in cells of the mammalian circadian clock. Nat Neurosci 3:1241–1247. doi:10.1038/81767
Ripperger JA, Schibler U (2006) Rhythmic CLOCK-BMAL1 binding to multiple E-box motifs drives circadian Dbp transcription and chromatin transitions. Nat Genet 38:369–374. doi:10.1038/ng1738
Etchegaray JP, Yang X, DeBruyne JP, Peters AH, Weaver DR, Jenuwein T, Reppert SM (2006) The polycomb group protein EZH2 is required for mammalian circadian clock function. J Biol Chem 281:21209–21215. doi:10.1074/jbc.M603722200
Sun Z, Feng D, Everett LJ, Bugge A, Lazar MA (2011) Circadian epigenomic remodeling and hepatic lipogenesis: lessons from HDAC3. Cold Spring Harb Sym 76:49. doi:10.1101/sqb.2011.76.011494
Azzi A, Dallmann R, Casserly A, Rehrauer H, Patrignani A, Maier B, Kramer A, Brown SA (2014) Circadian behavior is light-reprogrammed by plastic DNA methylation. Nat Neurosci 17:377–382. doi:10.1038/nn.3651
Menet JS, Pescatore S, Rosbash M (2014) CLOCK:BMAL1 is a pioneer-like transcription factor. Gene Dev 28:8–13. doi:10.1101/gad.228536.113
Etchegaray J, Lee C, Wade PA, Reppert SM (2003) Rhythmic histone acetylation underlies transcription in the mammalian circadian clock. Nature 421:177–182. doi:10.1038/nature01314
Curtis AM, Seo SP, Westgate EJ, Rudic RD, Smyth EM, Chakravarti D, FitzGerald GA, McNamara P (2004) Histone acetyltransferase-dependent chromatin remodeling and the vascular clock. J Biol Chem 279:7091–7097. doi:10.1074/jbc.M311973200
Cho H, Orphanides G, Sun X, Yang XJ, Ogryzko V, Lees E, Nakatani Y, Reinberg D (1998) A human RNA polymerase II complex containing factors that modify chromatin structure. Mol Cell Biol 18:5355–5363
Doi M, Hirayama J, Sassone-Corsi P (2006) Circadian regulator CLOCK is a histone acetyltransferase. Cell 125:497–508. doi:10.1016/j.cell.2006.03.033
Yamamoto T, Horikoshi M (1997) Novel substrate specificity of the histone acetyltransferase activity of HIV-1-Tat interactive protein Tip60. J Biol Chem 272:30595–30598. doi:10.1074/jbc.272.49.30595
Hirayama J, Sahar S, Grimaldi B, Tamaru T, Takamatsu K, Nakahata Y, Sassone-Corsi P (2007) CLOCK-mediated acetylation of BMAL1 controls circadian function. Nature 450:1086–1090. doi:10.1038/nature06394
DiTacchio L, Le HD, Vollmers C, Hatori M, Witcher M, Secombe J, Panda S (2011) Histone lysine demethylase JARID1a activates CLOCK-BMAL1 and influences the circadian clock. Science 333:1881–1885. doi:10.1126/science.1206022
Katada S, Sassone-Corsi P (2010) The histone methyltransferase MLL1 permits the oscillation of circadian gene expression. Nat Struct Mol Biol 17:1414–1421. doi:10.1038/nsmb.1961
Valekunja UK, Edgar RS, Oklejewicz M, van der Horst GTJ, O’Neill JS, Tamanini F, Turner DJ, Reddy AB (2013) Histone methyltransferase MLL3 contributes to genome-scale circadian transcription. Proc Natl Acad Sci USA 110:1554–1559. doi:10.1073/pnas.1214168110
Duong HA, Robles MS, Knutti D, Weitz CJ (2011) A molecular mechanism for circadian clock negative feedback. Science 332:1436–1439. doi:10.1126/science.1196766
Naruse Y, Oh-hashi K, Iijima N, Naruse M, Yoshioka H, Tanaka M (2004) Circadian and light-induced transcription of clock gene Per1 depends on histone acetylation and deacetylation. Mol Cell Biol 24:6278–6287. doi:10.1128/MCB.24.14.6278-6287.2004
Duong HA, Weitz CJ (2014) Temporal orchestration of repressive chromatin modifiers by circadian clock PERIOD complexes. Nat Struct Mol Biol 21:126–132. doi:10.1038/nsmb.2746
Brown SA, Ripperger J, Kadener S, Fleury-Olela F, Vilbois F, Rosbash M, Schibler U (2005) PERIOD1-associated proteins modulate the negative limb of the mammalian circadian oscillator. Science 308:693–696. doi:10.1126/science.1107373
Kim JY, Kwak PB, Weitz CJ (2014) Specificity in circadian clock feedback from targeted reconstitution of the NuRD corepressor. Mol Cell 56:738–748. doi:10.1016/j.molcel.2014.10.017
Nakahata Y, Kaluzova M, Grimaldi B, Sahar S, Hirayama J, Chen D, Guarente LP, Sassone-Corsi P (2008) The NAD+-dependent deacetylase SIRT1 modulates CLOCK-mediated chromatin remodeling and circadian control. Cell 134:329–340. doi:10.1016/j.cell.2008.07.002
Asher G, Gatfield D, Stratmann M, Reinke H, Dibner C, Kreppel F, Mostoslavsky R, Alt FW, Schibler U (2008) SIRT1 regulates circadian clock gene expression through PER2 deacetylation. Cell 134:317–328. doi:10.1016/j.cell.2008.06.050
Aguilar-Arnal L, Katada S, Orozco-Solis R, Sassone-Corsi P (2015) NAD+-SIRT1 control of H3K4 trimethylation through circadian deacetylation of MLL1. Nat Struct Mol Biol 22:312–318. doi:10.1038/nsmb.2990
Nakahata Y, Sahar S, Astarita G, Kaluzova M, Sassone-Corsi P (2009) Circadian control of the NAD+ salvage pathway by CLOCK-SIRT1. Science 324:654–657. doi:10.1126/science.1170803
Masri S, Rigor P, Cervantes M, Ceglia N, Sebastian C, Xiao C, Roqueta-Rivera M, Deng C, Osborne TF, Mostoslavsky R, Baldi P, Sassone-Corsi P (2014) Partitioning circadian transcription by SIRT6 leads to segregated control of cellular metabolism. Cell 158:659–672. doi:10.1016/j.cell.2014.06.050
Ginty DD, Kornhauser JM, Thompson MA, Bading H, Mayo KE, Takahashi JS, Greenberg ME (1993) Regulation of CREB phosphorylation in the suprachiasmatic nucleus by light and a circadian clock. Science 260:238–241. doi:10.1126/science.8097062
Travnickova-Bendova Z, Cermakian N, Reppert SM, Sassone-Corsi P (2002) Bimodal regulation of mPeriod promoters by CREB-dependent signaling and CLOCK/BMAL1 activity. Proc Natl Acad Sci USA 99:7728–7733. doi:10.1073/pnas.102075599
Tischkau SA, Mitchell JW, Tyan S-H, Buchanan GF, Gillette MU (2003) Ca2+/cAMP response element-binding protein (CREB)-dependent activation of Per1 is required for light-induced signaling in the suprachiasmatic nucleus circadian clock. J Biol Chem 278:718–723. doi:10.1074/jbc.M209241200
Lee B, Li A, Hansen KF, Cao R, Yoon JH, Obrietan K (2010) CREB influences timing and entrainment of the SCN circadian clock. J Biol Rhythms 25:410–420. doi:10.1177/0748730410381229
Sakamoto K, Norona FE, Alzate-Correa D, Scarberry D, Hoyt KR, Obrietan K (2013) Clock and light regulation of the CREB coactivator CRTC1 in the suprachiasmatic circadian clock. J Neurosci 33:9021–9027. doi:10.1523/JNEUROSCI.4202-12.2013
Jagannath A, Butler R, Godinho SI, Couch Y, Brown LA, Vasudevan SR, Flanagan KC, Anthony D, Churchill GC, Wood MJ, Steiner G, Ebeling M, Hossbach M, Wettstein JG, Duffield GE, Gatti S, Hankins MW, Foster RG, Peirson SN (2013) The CRTC1-SIK1 pathway regulates entrainment of the circadian clock. Cell 154:1100–1111. doi:10.1016/j.cell.2013.08.004
Sun X, Dang F, Zhang D, Yuan Y, Zhang C, Wu Y, Wang Y, Liu Y (2015) Glucagon-CREB/CRTC2 signaling cascade regulates hepatic BMAL1 protein. J Biol Chem 290:2189–2197. doi:10.1074/jbc.M114.612358
Koyanagi S, Hamdan AM, Horiguchi M, Kusunose N, Okamoto A, Matsunaga N, Ohdo S (2011) cAMP-response element (CRE)-mediated transcription by activating transcription factor-4 (ATF4) is essential for circadian expression of the Period2 gene. J Biol Chem 286:32416–32423. doi:10.1074/jbc.M111.258970
Tao W, Wu J, Zhang Q, Lai SS, Jiang S, Jiang C, Xu Y, Xue B, Du J, Li CJ (2015) EGR1 regulates hepatic clock gene amplitude by activating Per1 transcription. Sci Rep 5:15212. doi:10.1038/srep15212
Sakamoto KM, Fraser JK, Lee HJ, Lehman E, Gasson JC (1994) Granulocyte-macrophage colony-stimulating factor and interleukin-3 signaling pathways converge on the CREB-binding site in the human egr-1 promoter. Mol Cell Biol 14:5975–5985. doi:10.1128/MCB.14.9.5975
Kilduff TS, Vugrinic C, Lee SL, Milbrandt JD, Mikkelsen JD, O’Hara BF, Heller HC (1998) Characterization of the circadian system of NGFI-A and NGFI-A/NGFI-B deficient mice. J Biol Rhythms 13:347–357. doi:10.1177/074873098129000174
Kim SH, Yu HS, Park HG, Ahn YM, Kim YS, Lee YH, Ha K, Shin SY (2013) Egr1 regulates lithium-induced transcription of the Period 2 (PER2) gene. Biochim Biophys Acta 1832:1969–1979. doi:10.1016/j.bbadis.2013.06.010
Resuehr HE, Resuehr D, Olcese J (2009) Induction of mPer1 expression by GnRH in pituitary gonadotrope cells involves EGR-1. Mol Cell Endocrinol 311:120–125. doi:10.1016/j.mce.2009.07.005
Parsons MJ, Brancaccio M, Sethi S, Maywood ES, Satija R, Edwards JK, Jagannath A, Couch Y, Finelli MJ, Smyllie NJ, Esapa C, Butler R, Barnard AR, Chesham JE, Saito S, Joynson G, Wells S, Foster RG, Oliver PL, Simon MM, Mallon AM, Hastings MH, Nolan PM (2015) The regulatory factor ZFHX3 modifies circadian function in SCN via an AT motif-driven axis. Cell 162:607–621. doi:10.1016/j.cell.2015.06.060
Rossner MJ, Oster H, Wichert SP, Reinecke L, Wehr MC, Reinecke J, Eichele G, Taneja R, Nave KA (2008) Disturbed clockwork resetting in Sharp-1 and Sharp-2 single and double mutant mice. PLoS One 3:e2762. doi:10.1371/journal.pone.0002762
Honma S, Kawamoto T, Takagi Y, Fujimoto K, Sato F, Noshiro M, Kato Y, Honma KI (2002) Dec1 and Dec2 are regulators of the mammalian molecular clock. Nature 419:841–844. doi:10.1038/nature01123
Kawamoto T, Noshiro M, Sato F, Maemura K, Takeda N, Nagai R, Iwata T, Fujimoto K, Furukawa M, Miyazaki K, Honma S, Honma KI, Kato Y (2004) A novel autofeedback loop of Dec1 transcription involved in circadian rhythm regulation. Biochem Bioph Res Commun 313:117–124. doi:10.1016/j.bbrc.2003.11.099
Li Y, Song X, Ma Y, Liu J, Yang D, Yan B (2004) DNA binding, but not interaction with Bmal1, is responsible for DEC1-mediated transcription regulation of the circadian gene mPer1. Biochem J 382:895–904. doi:10.1042/BJ20040592
Cavadini G, Petrzilka S, Kohler P, Jud C, Tobler I, Birchler T, Fontana A (2007) TNF-alpha suppresses the expression of clock genes by interfering with E-box-mediated transcription. Proc Natl Acad Sci USA 104:12843–12848. doi:10.1073/pnas.0701466104
Bellet MM, Zocchi L, Sassone-Corsi P (2012) The RelB subunit of NFκB acts as a negative regulator of circadian gene expression. Cell Cycle 11:3304–3311. doi:10.4161/cc.21669
Ohno T, Onishi Y, Ishida N (2007) The negative transcription factor E4BP4 is associated with circadian clock protein PERIOD2. Biochem Bioph Res Commun 354:1010–1015. doi:10.1016/j.bbrc.2007.01.084
Ohno T, Onishi Y, Ishida N (2007) A novel E4BP4 element drives circadian expression of mPeriod2. Nucleic Acids Res 35:648–655. doi:10.1093/nar/gkl868
Tanoue S, Fujimoto K, Myung J, Hatanaka F, Kato Y, Takumi T (2015) DEC2-E4BP4 heterodimer represses the transcriptional enhancer activity of the EE element in the Per2 promoter. Front Neurol 6:166. doi:10.3389/fneur.2015.00166
Mitsui S, Yamaguchi S, Matsuo T, Ishida Y, Okamura H (2001) Antagonistic role of E4BP4 and PAR proteins in the circadian oscillatory mechanism. Genes Dev 15:995–1006. doi:10.1101/gad.873501
Ueda HR, Hayashi S, Chen W, Sano M, Machida M, Shigeyoshi Y, Iino M, Hashimoto S (2005) System-level identification of transcriptional circuits underlying mammalian circadian clocks. Nat Genet 37:187–192. doi:10.1038/ng1504
Miki T, Matsumoto T, Zhao Z, Lee CC (2013) p53 regulates Period2 expression and the circadian clock. Nat Commun 4:2444. doi:10.1038/ncomms3444
Gotoh T, Vila-Caballer M, Santos CS, Liu J, Yang J, Finkielstein CV (2014) The circadian factor Period 2 modulates p53 stability and transcriptional activity in unstressed cells. Mol Biol Cell 25:3081–3093. doi:10.1091/mbc.E14-05-0993
Gotoh T, Vila-Caballer M, Liu J, Schiffhauer S, Finkielstein CV (2015) Association of the circadian factor Period 2 to p53 influences p53’s function in DNA-damage signaling. Mol Biol Cell 26:359–372. doi:10.1091/mbc.E14-05-0994
Jiang W, Zhao S, Jiang X, Zhang E, Hu G, Hu B, Zheng P, Xiao J, Lu Z, Lu Y, Ni J, Chen C, Wang X, Yang L, Wan R (2016) The circadian clock gene Bmal1 acts as a potential anti-oncogene in pancreatic cancer by activating the p53 tumor suppressor pathway. Cancer Lett 371:314–325. doi:10.1016/j.canlet.2015.12.002
Miki T, Xu Z, Chen-Goodspeed M, Liu M, Van Oort-Jansen A, Rea MA, Zhao Z, Lee CC, Chang KS (2012) PML regulates PER2 nuclear localization and circadian function. EMBO J 31:1427–1439. doi:10.1038/emboj.2012.1
Fu L, Pelicano H, Liu J, Huang P, Lee C (2002) The circadian gene Period2 plays an important role in tumor suppression and DNA damage response in vivo. Cell 111:41–50. doi:10.1016/S0092-8674(02)00961-3
Altman BJ, Hsieh AL, Sengupta A, Krishnanaiah SY, Stine ZE, Walton ZE, Gouw AM, Venkataraman A, Li B, Goraksha-Hicks P, Diskin SJ, Bellovin DI, Simon MC, Rathmell JC, Lazar MA, Maris JM, Felsher DW, Hogenesch JB, Weljie AM, Dang CV (2015) MYC disrupts the circadian clock and metabolism in cancer cells. Cell Metab 22:1009–1019. doi:10.1016/j.cmet.2015.09.003
Benegiamo G, Brown SA, Panda S (2016) RNA dynamics in the control of circadian rhythm. Adv Exp Med Biol 907:107–122. doi:10.1007/978-3-319-29073-7_5
Preußner M, Heyd F (2016) Post-transcriptional control of the mammalian circadian clock: implications for health and disease. Pflug Arch Eur J Phy 468:983–991. doi:10.1007/s00424-016-1820-y
Chiang CK, Mehta N, Patel A, Zhang P, Ning Z, Mayne J, Sun WYL, Cheng HY, Figeys D (2014) The proteomic landscape of the suprachiasmatic nucleus clock reveals large-scale coordination of key biological processes. PLoS Genet 10:e1004695. doi:10.1371/journal.pgen.1004695
Mauvoisin D, Wang J, Jouffe C, Martin E, Atger F, Waridel P, Quadroni M, Gachon F, Naef F (2014) Circadian clock-dependent and -independent rhythmic proteomes implement distinct diurnal functions in mouse liver. Proc Natl Acad Sci USA 111:167–172. doi:10.1073/pnas.1314066111
Reddy AB, Karp NA, Maywood ES, Sage EA, Deery M, O’Neill JS, Wong GK, Chesham J, Odell M, Lilley KS, Kyriacou CP, Hastings MH (2006) Circadian orchestration of the hepatic proteome. Curr Biol 16:1107–1115. doi:10.1016/j.cub.2006.04.026
Elkon R, Ugalde AP, Agami R (2013) Alternative cleavage and polyadenylation: extent, regulation and function. Nat Rev Genet 14:496–506. doi:10.1038/nrg3482
Wang Y, Osterbur DL, Megaw PL, Tosini G, Fukuhara C, Green CB, Besharse JC (2001) Rhythmic expression of Nocturnin mRNA in multiple tissues of the mouse. BMC Dev Biol 1:9. doi:10.1186/1471-213X-1-9
Kojima S, Gendreau KL, Sher-Chen EL, Gao P, Green CB (2015) Changes in poly(A) tail length dynamics from the loss of the circadian deadenylase Nocturnin. Sci Rep 5:17059. doi:10.1038/srep17059
Robinson BG, Frim DM, Schwartz WJ, Majzoub JA (1988) Vasopressin mRNA in the suprachiasmatic nuclei: daily regulation of polyadenylate tail length. Science 241:342–344. doi:10.1126/science.3388044
Gerstner JR, Vanderheyden WM, LaVaute T, Westmark CJ, Rouhana L, Pack AI, Wickens M, Landry CF (2012) Time of day regulates subcellular trafficking, tripartite synaptic localization, and polyadenylation of the astrocytic Fabp7 mRNA. J Neurosci 32:1383–1394. doi:10.1523/JNEUROSCI.3228-11.2012
Villalba A, Coll O, Gebauer F (2011) Cytoplasmic polyadenylation and translational control. Curr Opin Genet Dev 21:452–457. doi:10.1016/j.gde.2011.04.006
Kojima S, Sher-Chen EL, Green CB (2012) Circadian control of mRNA polyadenylation dynamics regulates rhythmic protein expression. Genes Dev 26:2724–2736. doi:10.1101/gad.208306.112
Derti A, Garrett-Engele P, Macisaac KD, Stevens RC, Sriram S, Chen R, Rohl CA, Johnson JM, Babak T (2012) A quantitative atlas of polyadenylation in five mammals. Genome Res 22:1173–1183. doi:10.1101/gr.132563.111
Tian B, Hu J, Zhang H, Lutz CS (2005) A large-scale analysis of mRNA polyadenylation of human and mouse genes. Nucleic Acids Res 33:201–212. doi:10.1093/nar/gki158
Di Giammartino DC, Nishida K, Manley JL (2011) Mechanisms and consequences of alternative polyadenylation. Mol Cell 43:853–866. doi:10.1016/j.molcel.2011.08.017
Lai EC (2002) Micro RNAs are complementary to 3′ UTR sequence motifs that mediate negative post-transcriptional regulation. Nat Genet 30:363–364. doi:10.1038/ng865
Liu Y, Hu W, Murakawa Y, Yin J, Wang G, Landthaler M, Yan J (2013) Cold-induced RNA-binding proteins regulate circadian gene expression by controlling alternative polyadenylation. Sci Rep 3:2054. doi:10.1038/srep02054
Fustin JM, Doi M, Yamaguchi Y, Hida H, Nishimura S, Yoshida M, Isagawa T, Morioka MS, Kakeya H, Manabe I, Okamura H (2013) RNA-methylation-dependent RNA processing controls the speed of the circadian clock. Cell 155:793–806. doi:10.1016/j.cell.2013.10.026
Chen M, Manley JL (2009) Mechanisms of alternative splicing regulation: insights from molecular and genomics approaches. Nat Rev Mol Cell Biol 10:741–754. doi:10.1038/nrm2777
Majercak J, Chen WF, Edery I (2004) Splicing of the period gene 3′-terminal intron is regulated by light, circadian clock factors, and phospholipase C. Mol Cell Biol 24:3359–3372. doi:10.1128/MCB.24.8.3359-3372.2004
Low KH, Lim C, Ko HW, Edery I (2008) Natural variation in the splice site strength of a clock gene and species-specific thermal adaptation. Neuron 60:1054–1067. doi:10.1016/j.neuron.2008.10.048
Low KH, Chen WF, Yildirim E, Edery I (2012) Natural variation in the Drosophila melanogaster clock gene period modulates splicing of its 3′-terminal intron and mid-day siesta. PLoS One 7:e49536. doi:10.1371/journal.pone.0049536
Cheng Y, Gvakharia B, Hardin PE (1998) Two alternatively spliced transcripts from the Drosophila period gene rescue rhythms having different molecular and behavioral characteristics. Mol Cell Biol 18:6505–6514. doi:10.1128/MCB.18.11.6505
Majercak J, Sidote D, Hardin PE, Edery I (1999) How a circadian clock adapts to seasonal decreases in temperature and day length. Neuron 24:219–230. doi:10.1016/S0896-6273(00)80834-X
Sanchez SE, Petrillo E, Beckwith EJ, Zhang X, Rugnone ML, Hernando CE, Cuevas JC, Godoy Herz MA, Depetris-Chauvin A, Simpson CG, Brown JW, Cerdán PD, Borevitz JO, Mas P, Ceriani MF, Kornblihtt AR, Yanovsky MJ (2010) A methyl transferase links the circadian clock to the regulation of alternative splicing. Nature 468:112–116. doi:10.1038/nature09470
McGlincy NJ, Valomon A, Chesham JE, Maywood ES, Hastings MH, Ule J (2012) Regulation of alternative splicing by the circadian clock and food related cues. Genome Biol 13:R54. doi:10.1186/gb-2012-13-6-r54
Preußner M, Wilhelmi I, Schultz AS, Finkernagel F, Michel M, Möröy T, Heyd F (2014) Rhythmic U2af26 alternative splicing controls PERIOD1 stability and the circadian clock in mice. Mol Cell 54:651–662. doi:10.1016/j.molcel.2014.04.015
Morf J, Rey G, Schneider K, Stratmann M, Fujita J, Naef F, Schibler U (2012) Cold-inducible RNA-binding protein modulates circadian gene expression posttranscriptionally. Science 338:379–383. doi:10.1126/science.1217726
Zhang J, Fang Z, Jud C, Vansteensel MJ, Kaasik K, Lee CC, Albrecht U, Tamanini F, Meijer JH, Oostra BA, Nelson DL (2008) Fragile X-related proteins regulate mammalian circadian behavioral rhythms. Am J Hum Genet 83:43–52. doi:10.1016/j.ajhg.2008.06.003
Dockendorff TC, Su HS, McBride SM, Yang Z, Choi CH, Siwicki KK, Sehgal A, Jongens TA (2002) Drosophila lacking dfmr1 activity show defects in circadian output and fail to maintain courtship interest. Neuron 34:973–984. doi:10.1016/S0896-6273(02)00724-9
Kim M, Bellini M, Ceman S (2009) Fragile X mental retardation protein FMRP binds mRNAs in the nucleus. Mol Cell Biol 29:214–228. doi:10.1128/MCB.01377-08
Ji X, Kong J, Liebhaber SA (2011) An RNA-protein complex links enhanced nuclear 3′ processing with cytoplasmic mRNA stabilization. EMBO J 30:2622–2633. doi:10.1038/emboj.2011.171
Misquitta CM, Iyer VR, Werstiuk ES, Grover AK (2001) The role of 3′-untranslated region (3′-UTR) mediated mRNA stability in cardiovascular pathophysiology. Mol Cell Biochem 224:53–67. doi:10.1023/A:1011982932645
Woo KC, Ha DC, Lee KH, Kim DY, Kim TD, Kim KT (2010) Circadian amplitude of cryptochrome 1 is modulated by mRNA stability regulation via cytoplasmic hnRNP D oscillation. Mol Cell Biol 30:197–205. doi:10.1128/MCB.01154-09
Lee PT, Chao PK, Ou LC, Chuang JY, Lin YC, Chen SC, Chang HF, Law PY, Loh HH, Chao YS, Su TP, Yeh SH (2014) Morphine drives internal ribosome entry site-mediated hnRNP K translation in neurons through opioid receptor-dependent signaling. Nucleic Acids Res 42:13012–13025. doi:10.1093/nar/gku1016
Woo KC, Kim TD, Lee KH, Kim DY, Kim W, Lee KY, Kim KT (2009) Mouse period 2 mRNA circadian oscillation is modulated by PTB-mediated rhythmic mRNA degradation. Nucleic Acids Res 37:26–37. doi:10.1093/nar/gkn893
Kim SH, Lee KH, Kim DY, Kwak E, Kim S, Kim KT (2015) Rhythmic control of mRNA stability modulates circadian amplitude of mouse Period3 mRNA. J Neurochem 132:642–656. doi:10.1111/jnc.13027
Huang Y, Ainsley JA, Reijmers LG, Jackson FR (2013) Translational profiling of clock cells reveals circadianly synchronized protein synthesis. PLoS Biol 11:e1001703. doi:10.1371/journal.pbio.1001703
Huang Y, McNeil GP, Jackson FR (2014) Translational regulation of the DOUBLETIME/CKIδ/ε kinase by LARK contributes to circadian period modulation. PLoS Genet 10:e1004536. doi:10.1371/journal.pgen.1004536
Lim C, Lee J, Choi C, Kilman VL, Kim J, Park SM, Jang SK, Allada R, Choe J (2011) The novel gene twenty-four defines a critical translational step in the Drosophila clock. Nature 470:399–403. doi:10.1038/nature09728
Zhang Y, Ling J, Yuan C, Dubruille R, Emery P (2013) A role for Drosophila ATX2 in activation of PER translation and circadian behavior. Science 340:879–882. doi:10.1126/science.1234746
Lim C, Allada R (2013) ATAXIN-2 activates PERIOD translation to sustain circadian rhythms in Drosophila. Science 340:875–879. doi:10.1126/science.1234785
Jouffe C, Cretenet G, Symul L, Martin E, Atger F, Naef F, Gachon F (2013) The circadian clock coordinates ribosome biogenesis. PLoS Biol 11:e1001455. doi:10.1371/journal.pbio.1001455
Cao R, Lee B, Cho HY, Saklayen S, Obrietan K (2008) Photic regulation of the mTOR signaling pathway in the suprachiasmatic circadian clock. Mol Cell Neurosci 38:312–324. doi:10.1016/j.mcn.2008.03.005
Cao R, Anderson FE, Jung YJ, Dziema H, Obrietan K (2011) Circadian regulation of mammalian target of rapamycin signaling in the mouse suprachiasmatic nucleus. Neuroscience 181:79–88. doi:10.1016/j.neuroscience.2011.03.005
Cao R, Li A, Cho HY, Lee B, Obrietan K (2010) Mammalian target of rapamycin signaling modulates photic entrainment of the suprachiasmatic circadian clock. J Neurosci 30:6302–6314. doi:10.1523/JNEUROSCI.5482-09.2010
Zheng X, Sehgal A (2010) AKT and TOR signaling set the pace of the circadian pacemaker. Curr Biol 20:1203–1208. doi:10.1016/j.cub.2010.05.027
Cao R, Robinson B, Xu H, Gkogkas C, Khoutorsky A, Alain T, Yanagiya A, Nevarko T, Liu AC, Amir S, Sonenberg N (2013) Translational control of entrainment and synchrony of the suprachiasmatic circadian clock by mTOR/4E-BP1 signaling. Neuron 79:712–724. doi:10.1016/j.neuron.2013.06.026
Cao R, Gkogkas CG, de Zavalia N, Blum ID, Yanagiya A, Tsukumo Y, Xu H, Lee C, Storch KF, Liu AC, Amir S, Sonenberg N (2015) Light-regulated translational control of circadian behavior by eIF4E phosphorylation. Nat Neurosci 18:855–862. doi:10.1038/nn.4010
Butcher GQ, Lee B, Hsieh F, Obrietan K (2004) Light- and clock-dependent regulation of ribosomal S6 kinase activity in the suprachiasmatic nucleus. Eur J Neurosci 19:907–915. doi:10.1111/j.0953-816X.2004.03155.x
Tangredi MM, Ng FS, Jackson FR (2012) The C-terminal kinase and ERK-binding domains of Drosophila S6KII (RSK) are required for phosphorylation of the protein and modulation of circadian behavior. J Biol Chem 287:16748–16758. doi:10.1074/jbc.M111.315929
Akten B, Tangredi MM, Jauch E, Roberts MA, Ng F, Raabe T, Jackson FR (2009) Ribosomal s6 kinase cooperates with casein kinase 2 to modulate the Drosophila circadian molecular oscillator. J Neurosci 29:466–475. doi:10.1523/JNEUROSCI.4034-08.2009
Yates LA, Norbury CJ, Gilbert RJ (2013) The long and short of microRNA. Cell 153:516–519. doi:10.1016/j.cell.2013.04.003
Xu S, Witmer PD, Lumayag S, Kovacs B, Valle D (2007) MicroRNA (miRNA) transcriptome of mouse retina and identification of a sensory organ-specific miRNA cluster. J Biol Chem 282:25053–25066. doi:10.1074/jbc.M700501200
Na YJ, Sung JH, Lee SC, Lee YJ, Choi YJ, Park WY, Shin HS, Kim JH (2009) Comprehensive analysis of microRNA-mRNA co-expression in circadian rhythm. Exp Mol Med 41:638–647. doi:10.3858/emm.2009.41.9.070
Yang M, Lee JE, Padgett RW, Edery I (2008) Circadian regulation of a limited set of conserved microRNAs in Drosophila. BMC Genom 9:83. doi:10.1186/1471-2164-9-83
Kadener S, Menet JS, Sugino K, Horwich MD, Weissbein U, Nawathean P, Vagin VV, Zamore PD, Nelson SB, Rosbash M (2009) A role for microRNAs in the Drosophila circadian clock. Genes Dev 23:2179–2191. doi:10.1101/gad.1819509
Chen R, D’Alessandro M, Lee C (2013) miRNAs are required for generating a time delay critical for the circadian oscillator. Curr Biol 23:1959–1968. doi:10.1016/j.cub.2013.08.005
Du NH, Arpat AB, De Matos M, Gatfield D (2014) MicroRNAs shape circadian hepatic gene expression on a transcriptome-wide scale. Elife 3:e02510. doi:10.7554/eLife.02510
Cheng HY, Papp JW, Varlamova O, Dziema H, Russell B, Curfman JP, Nakazawa T, Shimizu K, Okamura H, Impey S, Obrietan K (2007) microRNA modulation of circadian-clock period and entrainment. Neuron 54:813–829. doi:10.1016/j.neuron.2007.05.017
Gatfield D, Le Martelot G, Vejnar CE, Gerlach D, Schaad O, Fleury-Olela F, Ruskeepää AL, Oresic M, Esau CC, Zdobnov EM, Schibler U (2009) Integration of microRNA miR-122 in hepatic circadian gene expression. Genes Dev 23:1313–1326. doi:10.1101/gad.1781009
Chen W, Liu Z, Li T, Zhang R, Xue Y, Zhong Y, Bai W, Zhou D, Zhao Z (2014) Regulation of Drosophila circadian rhythms by miRNA let-7 is mediated by a regulatory cycle. Nat Commun 5:5549. doi:10.1038/ncomms6549
Vodala S, Pescatore S, Rodriguez J, Buescher M, Chen YW, Weng R, Cohen SM, Rosbash M (2012) The oscillating miRNA 959-964 cluster impacts Drosophila feeding time and other circadian outputs. Cell Metab 16:601–612. doi:10.1016/j.cmet.2012.10.002
Luo W, Sehgal A (2012) Regulation of circadian behavioral output via a MicroRNA-JAK/STAT circuit. Cell 148:765–779. doi:10.1016/j.cell.2011.12.024
Zhang Y, Lamba P, Guo P, Emery P (2016) miR-124 regulates the phase of Drosophila circadian locomotor behavior. J Neurosci 36:2007–2013. doi:10.1523/JNEUROSCI.3286-15.2016
Shende VR, Goldrick MM, Ramani S, Earnest DJ (2011) Expression and rhythmic modulation of circulating microRNAs targeting the clock gene Bmal1 in mice. PLoS One 6:e22586. doi:10.1371/journal.pone.0022586
Tan X, Zhang P, Zhou L, Yin B, Pan H, Peng X (2012) Clock-controlled mir-142-3p can target its activator, Bmal1. BMC Mol Biol 13:27. doi:10.1186/1471-2199-13-27
Shende VR, Neuendorff N, Earnest DJ (2013) Role of miR-142-3p in the post-transcriptional regulation of the clock gene Bmal1 in the mouse SCN. PLoS One 8:e65300. doi:10.1371/journal.pone.0065300
Shende VR, Kim SM, Neuendorff N, Earnest DJ (2014) MicroRNAs function as cis- and trans-acting modulators of peripheral circadian clocks. FEBS Lett 588:3015–3022. doi:10.1016/j.febslet.2014.05.058
Nagel R, Clijsters L, Agami R (2009) The miRNA-192/194 cluster regulates the Period gene family and the circadian clock. FEBS J 276:5447–5455. doi:10.1111/j.1742-4658.2009.07229.x
Lee KH, Kim SH, Lee HR, Kim W, Kim DY, Shin JC, Yoo SH, Kim KT (2013) MicroRNA-185 oscillation controls circadian amplitude of mouse Cryptochrome 1 via translational regulation. Mol Biol Cell 24:2248–2255. doi:10.1091/mbc.E12-12-0849
Alvarez-Saavedra M, Antoun G, Yanagiya A, Oliva-Hernandez R, Cornejo-Palma D, Perez-Iratxeta C, Sonenberg N, Cheng HY (2011) miRNA-132 orchestrates chromatin remodeling and translational control of the circadian clock. Hum Mol Genet 20:731–751. doi:10.1093/hmg/ddq519
Querfurth C, Diernfellner AC, Gin E, Malzahn E, Höfer T, Brunner M (2011) Circadian conformational change of the Neurospora clock protein FREQUENCY triggered by clustered hyperphosphorylation of a basic domain. Mol Cell 43:713–722. doi:10.1016/j.molcel.2011.06.033
Curtin KD, Huang ZJ, Rosbash M (1995) Temporally regulated nuclear entry of the Drosophila period protein contributes to the circadian clock. Neuron 14:365–372. doi:10.1016/0896-6273(95)90292-9
Meyer P, Saez L, Young MW (2006) PER-TIM interactions in living Drosophila cells: an interval timer for the circadian clock. Science 311:226–229. doi:10.1126/science.1118126
Chang DC, Reppert SM (2003) A novel C-terminal domain of drosophila PERIOD inhibits dCLOCK:CYCLE-mediated transcription. Curr Biol 13:758–762. doi:10.1016/S0960-9822(03)00286-0
Saez L, Derasmo M, Meyer P, Stieglitz J, Young MW (2011) A key temporal delay in the circadian cycle of Drosophila is mediated by a nuclear localization signal in the timeless protein. Genetics 188:591–600. doi:10.1534/genetics.111.127225
Jang AR, Moravcevic K, Saez L, Young MW, Sehgal A (2015) Drosophila TIM binds importin α1, and acts as an adapter to transport PER to the nucleus. PLoS Genet 11:e1004974. doi:10.1371/journal.pgen.1004974
Lee Y, Jang AR, Francey LJ, Sehgal A, Hogenesch JB (2015) KPNB1 mediates PER/CRY nuclear translocation and circadian clock function. Elife 4:1–16. doi:10.7554/eLife.08647
Sakakida Y, Miyamoto Y, Nagoshi E, Akashi M, Nakamura TJ, Mamine T, Kasahara M, Minami Y, Yoneda Y, Takumi T (2005) Importin alpha/beta mediates nuclear transport of a mammalian circadian clock component, mCRY2, together with mPER2, through a bipartite nuclear localization signal. J Biol Chem 280:13272–13278. doi:10.1074/jbc.M413236200
Martinek S, Inonog S, Manoukian AS, Young MW (2001) A role for the segment polarity gene shaggy/GSK-3 in the Drosophila circadian clock. Cell 105:769–779. doi:10.1016/S0092-8674(01)00383-X
Iitaka C, Miyazaki K, Akaike T, Ishida N (2005) A role for glycogen synthase kinase-3beta in the mammalian circadian clock. J Biol Chem 280:29397–29402. doi:10.1074/jbc.M503526200
Lin J-M, Kilman VL, Keegan K, Paddock B, Emery-Le M, Rosbash M, Allada R (2002) A role for casein kinase 2alpha in the Drosophila circadian clock. Nature 420:816–820. doi:10.1038/nature01235
Akten B, Jauch E, Genova GK, Kim EY, Edery I, Raabe T, Jackson FR (2003) A role for CK2 in the Drosophila circadian oscillator. Nat Neurosci 6:251–257. doi:10.1038/nn1007
Top D, Harms E, Syed S, Adams EL, Saez L (2016) GSK-3 and CK2 kinases converge on timeless to regulate the master clock. Cell Rep 16:357–367. doi:10.1016/j.celrep.2016.06.005
Ko HW, Kim EY, Chiu J, Vanselow JT, Kramer A, Edery I (2010) A hierarchical phosphorylation cascade that regulates the timing of PERIOD nuclear entry reveals novel roles for proline-directed kinases and GSK-3beta/SGG in circadian clocks. J Neurosci 30:12664–12675. doi:10.1523/JNEUROSCI.1586-10.2010
Lin J-M, Schroeder A, Allada R (2005) In vivo circadian function of casein kinase 2 phosphorylation sites in Drosophila PERIOD. J Neurosci 25:11175–11183. doi:10.1523/JNEUROSCI.2159-05.2005
Dusik V, Senthilan PR, Mentzel B, Hartlieb H, Wülbeck C, Yoshii T, Raabe T, Helfrich-Förster C (2014) The MAP kinase p38 is part of Drosophila melanogaster’s circadian clock. PLoS Genet 10:e1004565. doi:10.1371/journal.pgen.1004565
Kim EY, Jeong EH, Park S, Jeong H-J, Edery I, Cho JW (2012) A role for O-GlcNAcylation in setting circadian clock speed. Genes Dev 26:490–502. doi:10.1101/gad.182378.111
Takano A, Isojima Y, Nagai K (2004) Identification of mPer1 phosphorylation sites responsible for the nuclear entry. J Biol Chem 279:32578–32585. doi:10.1074/jbc.M403433200
Mehta N, Cheng AH, Chiang C-K, Mendoza-Viveros L, Ling HH, Patel A, Xu B, Figeys D, Cheng H-YM (2015) GRK2 fine-tunes circadian clock speed and entrainment via transcriptional and post-translational control of PERIOD proteins. Cell Rep 12:1272–1288. doi:10.1016/j.celrep.2015.07.037
Kurabayashi N, Hirota T, Sakai M, Sanada K, Fukada Y (2010) DYRK1A and glycogen synthase kinase 3beta, a dual-kinase mechanism directing proteasomal degradation of CRY2 for circadian timekeeping. Mol Cell Biol 30:1757–1768. doi:10.1128/MCB.01047-09
Gallego M, Virshup DM (2007) Post-translational modifications regulate the ticking of the circadian clock. Nat Rev Mol Cell Biol 8:139–148. doi:10.1038/nrm2106
Glickman MH, Ciechanover A (2002) The ubiquitin-proteasome proteolytic pathway: destruction for the sake of construction. Physiol Rev 82:373–428. doi:10.1152/physrev.00027.2001
Godinho SIH, Maywood ES, Shaw L, Tucci V, Barnard AR, Busino L, Pagano M, Kendall R, Quwailid MM, Romero MR, O’neill J, Chesham JE, Brooker D, Lalanne Z, Hastings MH, Nolan PM (2007) The after-hours mutant reveals a role for Fbxl3 in determining mammalian circadian period. Science 316:897–900. doi:10.1126/science.1141138
Siepka SM, Yoo S-H, Park J, Song W, Kumar V, Hu Y, Lee C, Takahashi JS (2007) Circadian mutant Overtime reveals F-box protein FBXL3 regulation of cryptochrome and period gene expression. Cell 129:1011–1023. doi:10.1016/j.cell.2007.04.030
Busino L, Bassermann F, Maiolica A, Lee C, Nolan PM, Godinho SIH, Draetta GF, Pagano M (2007) SCFFbxl3 controls the oscillation of the circadian clock by directing the degradation of cryptochrome proteins. Science 316:900–904. doi:10.1126/science.1141194
Hirano A, Yumimoto K, Tsunematsu R, Matsumoto M, Oyama M, Kozuka-Hata H, Nakagawa T, Lanjakornsiripan D, Nakayama KI, Fukada Y (2013) FBXL21 regulates oscillation of the circadian clock through ubiquitination and stabilization of cryptochromes. Cell 152:1106–1118. doi:10.1016/j.cell.2013.01.054
Yoo S-H, Mohawk JA, Siepka SM, Shan Y, Huh SK, Hong H-K, Kornblum I, Kumar V, Koike N, Xu M, Nussbaum J, Liu X, Chen Z, Chen ZJ, Green CB, Takahashi JS (2013) Competing E3 ubiquitin ligases govern circadian periodicity by degradation of CRY in nucleus and cytoplasm. Cell 152:1091–1105. doi:10.1016/j.cell.2013.01.055
Lamia KA, Sachdeva UM, DiTacchio L, Williams EC, Alvarez JG, Egan DF, Vasquez DS, Juguilon H, Panda S, Shaw RJ, Thompson CB, Evans RM (2009) AMPK regulates the circadian clock by cryptochrome phosphorylation and degradation. Science 326:437–440. doi:10.1126/science.1172156
Gao P, Yoo S-H, Lee K-J, Rosensweig C, Takahashi JS, Chen BP, Green CB (2013) Phosphorylation of the cryptochrome 1 C-terminal tail regulates circadian period length. J Biol Chem 288:35277–35286. doi:10.1074/jbc.M113.509604
Tong X, Buelow K, Guha A, Rausch R, Yin L (2012) USP2a protein deubiquitinates and stabilizes the circadian protein CRY1 in response to inflammatory signals. J Biol Chem 287:25280–25291. doi:10.1074/jbc.M112.340786
Ohsaki K, Oishi K, Kozono Y, Nakayama K, Nakayama KI, Ishida N (2008) The role of β-TrCP1 and β-TrCP2 in circadian rhythm generation by mediating degradation of clock protein PER2. J Biochem 144:609–618. doi:10.1093/jb/mvn112
Eide EJ, Woolf MF, Kang H, Woolf P, Hurst W, Camacho F, Vielhaber EL, Giovanni A, Virshup DM (2005) Control of mammalian circadian rhythm by CKIepsilon-regulated proteasome-mediated PER2 degradation. Mol Cell Biol 25:2795–2807. doi:10.1128/MCB.25.7.2795-2807.2005
Shirogane T, Jin J, Ang XL, Harper JW (2005) SCFbeta-TRCP controls clock-dependent transcription via casein kinase 1-dependent degradation of the mammalian period-1 (Per1) protein. J Biol Chem 280:26863–26872. doi:10.1074/jbc.M502862200
Grima B, Lamouroux A, Chélot E, Papin C, Limbourg-Bouchon B, Rouyer F (2002) The F-box protein slimb controls the levels of clock proteins period and timeless. Nature 420:178–182. doi:10.1038/nature01122
Ko HW, Jiang J, Edery I (2002) Role for Slimb in the degradation of Drosophila Period protein phosphorylated by Doubletime. Nature 420:673–678. doi:10.1038/nature01272
Meng Q-J, Logunova L, Maywood ES, Gallego M, Lebiecki J, Brown TM, Sládek M, Semikhodskii AS, Glossop NRJ, Piggins HD, Chesham JE, Bechtold DA, Yoo S-H, Takahashi JS, Virshup DM, Boot-Handford RP, Hastings MH, Loudon ASI (2008) Setting clock speed in mammals: the CK1 epsilon tau mutation in mice accelerates circadian pacemakers by selectively destabilizing PERIOD proteins. Neuron 58:78–88. doi:10.1016/j.neuron.2008.01.019
Price JL, Blau J, Rothenfluh A, Abodeely M, Kloss B, Young MW (1998) double-time is a novel Drosophila clock gene that regulates PERIOD protein accumulation. Cell 94:83–95. doi:10.1016/S0092-8674(00)81224-6
Chiu JC, Vanselow JT, Kramer A, Edery I (2008) The phospho-occupancy of an atypical SLIMB-binding site on PERIOD that is phosphorylated by DOUBLETIME controls the pace of the clock. Genes Dev 22:1758–1772. doi:10.1101/gad.1682708
Syed S, Saez L, Young MW (2011) Kinetics of doubletime kinase-dependent degradation of the Drosophila period protein. J Biol Chem 286:27654–27662. doi:10.1074/jbc.M111.243618
Shanware NP, Hutchinson JA, Kim SH, Zhan L, Bowler MJ, Tibbetts RS (2011) Casein kinase 1-dependent phosphorylation of familial advanced sleep phase syndrome-associated residues controls PERIOD 2 stability. J Biol Chem 286:12766–12774. doi:10.1074/jbc.M111.224014
Toh KL, Jones CR, He Y, Eide EJ, Hinz WA, Virshup DM, Ptácek LJ, Fu YH (2001) An hPer2 phosphorylation site mutation in familial advanced sleep phase syndrome. Science 291:1040–1043. doi:10.1126/science.1057499
Lee B, Almad A, Butcher GQ, Obrietan K (2007) Protein kinase C modulates the phase-delaying effects of light in the mammalian circadian clock. Eur J Neurosci 26:451–462. doi:10.1111/j.1460-9568.2007.05664.x
Jakubcakova V, Oster H, Tamanini F, Cadenas C, Leitges M, van der Horst GTJ, Eichele G (2007) Light entrainment of the mammalian circadian clock by a PRKCA-dependent posttranslational mechanism. Neuron 54:831–843. doi:10.1016/j.neuron.2007.04.031
Lee H, Chen R, Kim H, Etchegaray J-P, Weaver DR, Lee C (2011) The period of the circadian oscillator is primarily determined by the balance between casein kinase 1 and protein phosphatase 1. Proc Natl Acad Sci USA 108:16451–16456. doi:10.1073/pnas.1107178108
Fang Y, Sathyanarayanan S, Sehgal A (2007) Post-translational regulation of the Drosophila circadian clock requires protein phosphatase 1 (PP1). Genes Dev 21:1506–1518. doi:10.1101/gad.1541607
Gallego M, Kang H, Virshup DM (2006) Protein phosphatase 1 regulates the stability of the circadian protein PER2. Biochem J 399:169–175. doi:10.1042/BJ20060678
Sathyanarayanan S, Zheng X, Xiao R, Sehgal A (2004) Posttranslational regulation of Drosophila PERIOD protein by protein phosphatase 2A. Cell 116:603–615. doi:10.1016/S0092-8674(04)00128-X
Lee J, Lee Y, Lee MJ, Park E, Kang SH, Chung CH, Lee KH, Kim K (2008) Dual modification of BMAL1 by SUMO2/3 and ubiquitin promotes circadian activation of the CLOCK/BMAL1 complex. Mol Cell Biol 28:6056–6065. doi:10.1128/MCB.00583-08
Cardone L, Hirayama J, Giordano F, Tamaru T, Palvimo JJ, Sassone-Corsi P (2005) Circadian clock control by SUMOylation of BMAL1. Science 309:1390–1394. doi:10.1126/science.1110689
Sahar S, Zocchi L, Kinoshita C, Borrelli E, Sassone-Corsi P (2010) Regulation of BMAL1 protein stability and circadian function by GSK3beta-mediated phosphorylation. PLoS ONE 5:e8561. doi:10.1371/journal.pone.0008561
Zhang L, Abraham D, Lin S-T, Oster H, Eichele G, Fu Y-H, Ptáček LJ (2012) PKCγ participates in food entrainment by regulating BMAL1. Proc Natl Acad Sci USA 109:20679–20684. doi:10.1073/pnas.1218699110
Li M-D, Ruan H-B, Hughes ME, Lee J-S, Singh JP, Jones SP, Nitabach MN, Yang X (2013) O-GlcNAc signaling entrains the circadian clock by inhibiting BMAL1/CLOCK ubiquitination. Cell Metab 17:303–310. doi:10.1016/j.cmet.2012.12.015
Ma Y-T, Luo H, Guan W-J, Zhang H, Chen C, Wang Z, Li J-D (2013) O-GlcNAcylation of BMAL1 regulates circadian rhythms in NIH3T3 fibroblasts. Biochem Biophys Res Commun 431:382–387. doi:10.1016/j.bbrc.2013.01.043
Gossan NC, Zhang F, Guo B, Jin D, Yoshitane H, Yao A, Glossop N, Zhang YQ, Fukada Y, Meng Q-J (2014) The E3 ubiquitin ligase UBE3A is an integral component of the molecular circadian clock through regulating the BMAL1 transcription factor. Nucleic Acids Res 42:5765–5775. doi:10.1093/nar/gku225
Scoma HD, Humby M, Yadav G, Zhang Q, Fogerty J, Besharse JC (2011) The de-ubiquitinylating enzyme, USP2, is associated with the circadian clockwork and regulates its sensitivity to light. PLoS One 6:e25382. doi:10.1371/journal.pone.0025382
Lamaze A, Lamouroux A, Vias C, Hung H-C, Weber F, Rouyer F (2011) The E3 ubiquitin ligase CTRIP controls CLOCK levels and PERIOD oscillations in Drosophila. EMBO Rep 12:549–557. doi:10.1038/embor.2011.64
Luo W, Li Y, Tang C-HA, Abruzzi KC, Rodriguez J, Pescatore S, Rosbash M (2012) CLOCK deubiquitylation by USP8 inhibits CLK/CYC transcription in Drosophila. Genes Dev 26:2536–2549. doi:10.1101/gad.200584.112
Spengler ML, Kuropatwinski KK, Schumer M, Antoch MP (2009) A serine cluster mediates BMAL1-dependent CLOCK phosphorylation and degradation. Cell Cycle 8:4138–4146. doi:10.4161/cc.8.24.10273
Yu W, Zheng H, Price JL, Hardin PE (2009) DOUBLETIME plays a noncatalytic role to mediate CLOCK phosphorylation and repress CLOCK-dependent transcription within the Drosophila circadian clock. Mol Cell Biol 29:1452–1458. doi:10.1128/MCB.01777-08
Yu W, Zheng H, Houl JH, Dauwalder B, Hardin PE (2006) PER-dependent rhythms in CLK phosphorylation and E-box binding regulate circadian transcription. Genes Dev 20:723–733. doi:10.1101/gad.1404406
Kim EY, Edery I (2006) Balance between DBT/CKIepsilon kinase and protein phosphatase activities regulate phosphorylation and stability of Drosophila CLOCK protein. Proc Natl Acad Sci USA 103:6178–6183. doi:10.1073/pnas.0511215103
Szabó A, Papin C, Zorn D, Ponien P, Weber F, Raabe T, Rouyer F (2013) The CK2 kinase stabilizes CLOCK and represses its activity in the Drosophila circadian oscillator. PLoS Biol 11:e1001645. doi:10.1371/journal.pbio.1001645
Andreazza S, Bouleau S, Martin B, Lamouroux A, Ponien P, Papin C, Chélot E, Jacquet E, Rouyer F (2015) Daytime CLOCK dephosphorylation is controlled by STRIPAK complexes in Drosophila. Cell Rep 11:1266–1279. doi:10.1016/j.celrep.2015.04.033
Koh K, Zheng X, Sehgal A (2006) JETLAG resets the Drosophila circadian clock by promoting light-induced degradation of TIMELESS. Science 312:1809–1812. doi:10.1126/science.1124951
Stoleru D, Nawathean P, Fernández M de la P, Menet JS, Ceriani MF, Rosbash M (2007) The Drosophila circadian network is a seasonal timer. Cell 129:207–219. doi:10.1016/j.cell.2007.02.038
Peschel N, Chen KF, Szabo G, Stanewsky R (2009) Light-dependent interactions between the Drosophila circadian clock factors cryptochrome, jetlag, and timeless. Curr Biol 19:241–247. doi:10.1016/j.cub.2008.12.042
Knowles A, Koh K, Wu J-T, Chien C-T, Chamovitz DA, Blau J (2009) The COP9 signalosome is required for light-dependent timeless degradation and Drosophila clock resetting. J Neurosci 29:1152–1162. doi:10.1523/JNEUROSCI.0429-08.2009
Acknowledgments
This work was supported by grants to H.-Y.M.C. from the Canadian Institute of Health Research (CIHR) and the Natural Sciences and Engineering Research Council (NSERC) of Canada. H.-Y.M.C. is a Tier II Canada Research Chair (CRC) in Molecular Genetics of Biological Clocks. L.M.-V. is supported by a graduate scholarship from the Consejo Nacional de Ciencia y Tecnologia (CONACYT) of Mexico. P.B.-C., S.H. and A.H.C. are supported by NSERC Post-Graduate Scholarships. A.H.C. was previously supported by a CIHR-funded training fellowship from Sleep and Biological Rhythms Toronto. This work is dedicated to the memory of Harrod Ho Pak Ling, enthusiast of all things rhythmic.
Author information
Authors and Affiliations
Corresponding author
Additional information
L. Mendoza-Viveros, P. Bouchard-Cannon and S. Hegazi contributed equally.
Rights and permissions
About this article
Cite this article
Mendoza-Viveros, L., Bouchard-Cannon, P., Hegazi, S. et al. Molecular modulators of the circadian clock: lessons from flies and mice. Cell. Mol. Life Sci. 74, 1035–1059 (2017). https://doi.org/10.1007/s00018-016-2378-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-016-2378-8