Abstract
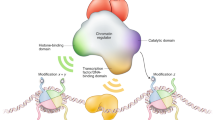
More and more studies have shown chromatin remodelers and histone modifiers play essential roles in regulating developmental patterns by organizing specific chromosomal architecture to establish programmed transcriptional profiles, with implications that histone chaperones execute a coordinating role in these processes. Chromatin assembly factor-1 (CAF-1), an evolutionarily conserved three-subunit protein complex, was identified as a histone chaperone coupled with DNA replication and repair in cultured mammalian cells and yeasts. Interestingly, recent findings indicate CAF-1 may have important regulatory roles during development by interacting with specific transcription factors and epigenetic regulators. In this review, we focus on the essential roles of CAF-1 in regulating heterochromatin organization, asymmetric cell division, and specific signal transduction through epigenetic modulations of the chromatin. In the end, we aim at providing a current image of facets of CAF-1 as a histone chaperone to orchestrate cell proliferation and differentiation during multi-cellular organism development.


Similar content being viewed by others
References
Eccleston A, Cesari F, Skipper M (2013) Transcription and epigenetics. Nature 502(7472):461. doi:10.1038/502461a
Landry JW, Banerjee S, Taylor B, Aplan PD, Singer A, Wu C (2011) Chromatin remodeling complex NURF regulates thymocyte maturation. Genes Dev 25(3):275–286. doi:10.1101/gad.2007311
Song H, Spichiger-Haeusermann C, Basler K (2009) The ISWI-containing NURF complex regulates the output of the canonical Wingless pathway. EMBO Rep 10(10):1140–1146. doi:10.1038/embor.2009.157
Saitou M, Kagiwada S, Kurimoto K (2012) Epigenetic reprogramming in mouse pre-implantation development and primordial germ cells. Development 139(1):15–31. doi:10.1242/dev.050849
Cantone I, Fisher AG (2013) Epigenetic programming and reprogramming during development. Nat Struct Mol Biol 20(3):282–289. doi:10.1038/nsmb.2489
Moshkin YM, Kan TW, Goodfellow H, Bezstarosti K, Maeda RK, Pilyugin M, Karch F, Bray SJ, Demmers JA, Verrijzer CP (2009) Histone chaperones ASF1 and NAP1 differentially modulate removal of active histone marks by LID-RPD3 complexes during NOTCH silencing. Mol Cell 35(6):782–793. doi:10.1016/j.molcel.2009.07.020
Goodfellow H, Krejci A, Moshkin Y, Verrijzer CP, Karch F, Bray SJ (2007) Gene-specific targeting of the histone chaperone asf1 to mediate silencing. Dev Cell 13(4):593–600. doi:10.1016/j.devcel.2007.08.021
Jullien J, Astrand C, Szenker E, Garrett N, Almouzni G, Gurdon JB (2012) HIRA dependent H3.3 deposition is required for transcriptional reprogramming following nuclear transfer to Xenopus oocytes. Epigenet Chromatin 5(1):17. doi:10.1186/1756-8935-5-17
Szenker E, Lacoste N, Almouzni G (2012) A developmental requirement for HIRA-dependent H3.3 deposition revealed at gastrulation in Xenopus. Cell reports 1(6):730–740. doi:10.1016/j.celrep.2012.05.006
Hajkova P, Ancelin K, Waldmann T, Lacoste N, Lange UC, Cesari F, Lee C, Almouzni G, Schneider R, Surani MA (2008) Chromatin dynamics during epigenetic reprogramming in the mouse germ line. Nature 452(7189):877–881. doi:10.1038/nature06714
Smith S, Stillman B (1989) Purification and characterization of CAF-I, a human cell factor required for chromatin assembly during DNA replication in vitro. Cell 58(1):15–25
Quivy JP, Gerard A, Cook AJ, Roche D, Almouzni G (2008) The HP1-p150/CAF-1 interaction is required for pericentric heterochromatin replication and S-phase progression in mouse cells. Nat Struct Mol Biol 15(9):972–979
Polo SE, Roche D, Almouzni G (2006) New histone incorporation marks sites of UV repair in human cells. Cell 127(3):481–493. doi:10.1016/j.cell.2006.08.049
Ye X, Franco AA, Santos H, Nelson DM, Kaufman PD, Adams PD (2003) Defective S phase chromatin assembly causes DNA damage, activation of the S phase checkpoint, and S phase arrest. Mol Cell 11(2):341–351
Baldeyron C, Soria G, Roche D, Cook AJ, Almouzni G (2011) HP1alpha recruitment to DNA damage by p150CAF-1 promotes homologous recombination repair. J Cell Biol 193(1):81–95. doi:10.1083/jcb.201101030
Quivy JP, Roche D, Kirschner D, Tagami H, Nakatani Y, Almouzni G (2004) A CAF-1 dependent pool of HP1 during heterochromatin duplication. EMBO J 23(17):3516–3526. doi:10.1038/sj.emboj.7600362
Adam S, Polo SE, Almouzni G (2013) Transcription recovery after DNA damage requires chromatin priming by the H3.3 histone chaperone HIRA. Cell 155(1):94–106. doi:10.1016/j.cell.2013.08.029
Nakano S, Stillman B, Horvitz HR (2011) Replication-coupled chromatin assembly generates a neuronal bilateral asymmetry in C. elegans. Cell 147(7):1525–1536. doi:10.1016/j.cell.2011.11.053
Yu Z, Wu H, Chen H, Wang R, Liang X, Liu J, Li C, Deng WM, Jiao R (2013) CAF-1 promotes Notch signaling through epigenetic control of target gene expression during Drosophila development. Development 140(17):3635–3644. doi:10.1242/dev.094599
Huang H, Yu Z, Zhang S, Liang X, Chen J, Li C, Ma J, Jiao R (2010) Drosophila CAF-1 regulates HP1-mediated epigenetic silencing and pericentric heterochromatin stability. J Cell Sci 123(Pt 16):2853–2861. doi:10.1242/jcs.063610
Zeng A, Li YQ, Wang C, Han XS, Li G, Wang JY, Li DS, Qin YW, Shi Y, Brewer G, Jing Q (2013) Heterochromatin protein 1 promotes self-renewal and triggers regenerative proliferation in adult stem cells. J Cell Biol 201(3):409–425. doi:10.1083/jcb.201207172
Groth A, Rocha W, Verreault A, Almouzni G (2007) Chromatin challenges during DNA replication and repair. Cell 128(4):721–733. doi:10.1016/j.cell.2007.01.030
Li Q, Zhou H, Wurtele H, Davies B, Horazdovsky B, Verreault A, Zhang Z (2008) Acetylation of histone H3 lysine 56 regulates replication-coupled nucleosome assembly. Cell 134(2):244–255. doi:10.1016/j.cell.2008.06.018
Das C, Lucia MS, Hansen KC, Tyler JK (2009) CBP/p300-mediated acetylation of histone H3 on lysine 56. Nature 459(7243):113–117. doi:10.1038/nature07861
Tjeertes JV, Miller KM, Jackson SP (2009) Screen for DNA-damage-responsive histone modifications identifies H3K9Ac and H3K56Ac in human cells. EMBO J 28(13):1878–1889. doi:10.1038/emboj.2009.119
Vempati RK, Jayani RS, Notani D, Sengupta A, Galande S, Haldar D (2010) p300-mediated acetylation of histone H3 lysine 56 functions in DNA damage response in mammals. J Biol Chem 285(37):28553–28564. doi:10.1074/jbc.M110.149393
Burgess RJ, Zhou H, Han J, Zhang Z (2010) A role for Gcn5 in replication-coupled nucleosome assembly. Mol Cell 37(4):469–480. doi:10.1016/j.molcel.2010.01.020
Allis CD, Chicoine LG, Richman R, Schulman IG (1985) Deposition-related histone acetylation in micronuclei of conjugating Tetrahymena. Proc Natl Acad Sci USA 82(23):8048–8052
Sobel RE, Cook RG, Allis CD (1994) Non-random acetylation of histone H4 by a cytoplasmic histone acetyltransferase as determined by novel methodology. J Biol Chem 269(28):18576–18582
Sobel RE, Cook RG, Perry CA, Annunziato AT, Allis CD (1995) Conservation of deposition-related acetylation sites in newly synthesized histones H3 and H4. Proc Natl Acad Sci USA 92(4):1237–1241
Verreault A, Kaufman PD, Kobayashi R, Stillman B (1996) Nucleosome assembly by a complex of CAF-1 and acetylated histones H3/H4. Cell 87(1):95–104
Zhang H, Han J, Kang B, Burgess R, Zhang Z (2012) Human histone acetyltransferase 1 protein preferentially acetylates H4 histone molecules in H3.1-H4 over H3.3-H4. J Biol Chem 287(9):6573–6581. doi:10.1074/jbc.M111.312637
Ma XJ, Wu J, Altheim BA, Schultz MC, Grunstein M (1998) Deposition-related sites K5/K12 in histone H4 are not required for nucleosome deposition in yeast. Proc Natl Acad Sci USA 95(12):6693–6698
Mello JA, Sillje HH, Roche DM, Kirschner DB, Nigg EA, Almouzni G (2002) Human Asf1 and CAF-1 interact and synergize in a repair-coupled nucleosome assembly pathway. EMBO Rep 3(4):329–334. doi:10.1093/embo-reports/kvf068
Shibahara K, Stillman B (1999) Replication-dependent marking of DNA by PCNA facilitates CAF-1-coupled inheritance of chromatin. Cell 96(4):575–585
Winkler DD, Zhou H, Dar MA, Zhang Z, Luger K (2012) Yeast CAF-1 assembles histone (H3-H4)2 tetramers prior to DNA deposition. Nucleic Acids Res 40(20):10139–10149. doi:10.1093/nar/gks812
Maison C, Almouzni G (2004) HP1 and the dynamics of heterochromatin maintenance. Nat Rev Mol Cell Biol 5(4):296–304. doi:10.1038/nrm1355
Huang S, Zhou H, Tarara J, Zhang Z (2007) A novel role for histone chaperones CAF-1 and Rtt106p in heterochromatin silencing. EMBO J 26(9):2274–2283. doi:10.1038/sj.emboj.7601670
Enomoto S, Berman J (1998) Chromatin assembly factor I contributes to the maintenance, but not the re-establishment, of silencing at the yeast silent mating loci. Genes Dev 12(2):219–232
Enomoto S, McCune-Zierath PD, Gerami-Nejad M, Sanders MA, Berman J (1997) RLF2, a subunit of yeast chromatin assembly factor-I, is required for telomeric chromatin function in vivo. Genes Dev 11(3):358–370
Kaufman PD, Kobayashi R, Stillman B (1997) Ultraviolet radiation sensitivity and reduction of telomeric silencing in Saccharomyces cerevisiae cells lacking chromatin assembly factor-I. Genes Dev 11(3):345–357
Houlard M, Berlivet S, Probst AV, Quivy JP, Hery P, Almouzni G, Gerard M (2006) CAF-1 is essential for heterochromatin organization in pluripotent embryonic cells. PLoS Genet 2(11):e181. doi:10.1371/journal.pgen.0020181
Loyola A, Tagami H, Bonaldi T, Roche D, Quivy JP, Imhof A, Nakatani Y, Dent SY, Almouzni G (2009) The HP1alpha-CAF1-SetDB1-containing complex provides H3K9me1 for Suv39-mediated K9me3 in pericentric heterochromatin. EMBO Rep 10(7):769–775. doi:10.1038/embor.2009.90
Ciccia A, Elledge SJ (2010) The DNA damage response: making it safe to play with knives. Mol Cell 40(2):179–204. doi:10.1016/j.molcel.2010.09.019
Tamburini BA, Tyler JK (2005) Localized histone acetylation and deacetylation triggered by the homologous recombination pathway of double-strand DNA repair. Mol Cell Biol 25(12):4903–4913. doi:10.1128/MCB.25.12.4903-4913.2005
Papamichos-Chronakis M, Krebs JE, Peterson CL (2006) Interplay between Ino80 and Swr1 chromatin remodeling enzymes regulates cell cycle checkpoint adaptation in response to DNA damage. Genes Dev 20(17):2437–2449. doi:10.1101/gad.1440206
Gaillard PH, Martini EM, Kaufman PD, Stillman B, Moustacchi E, Almouzni G (1996) Chromatin assembly coupled to DNA repair: a new role for chromatin assembly factor I. Cell 86(6):887–896
Nabatiyan A, Szuts D, Krude T (2006) Induction of CAF-1 expression in response to DNA strand breaks in quiescent human cells. Mol Cell Biol 26(5):1839–1849. doi:10.1128/MCB.26.5.1839-1849.2006
Jiao R, Harrigan JA, Shevelev I, Dietschy T, Selak N, Indig FE, Piotrowski J, Janscak P, Bohr VA, Stagljar I (2007) The Werner syndrome protein is required for recruitment of chromatin assembly factor 1 following DNA damage. Oncogene 26(26):3811–3822. doi:10.1038/sj.onc.1210150
Jiao R, Bachrati CZ, Pedrazzi G, Kuster P, Petkovic M, Li JL, Egli D, Hickson ID, Stagljar I (2004) Physical and functional interaction between the Bloom’s syndrome gene product and the largest subunit of chromatin assembly factor 1. Mol Cell Biol 24(11):4710–4719. doi:10.1128/MCB.24.11.4710-4719.2004
Fussner E, Djuric U, Strauss M, Hotta A, Perez-Iratxeta C, Lanner F, Dilworth FJ, Ellis J, Bazett-Jones DP (2011) Constitutive heterochromatin reorganization during somatic cell reprogramming. EMBO J 30(9):1778–1789. doi:10.1038/emboj.2011.96
Shermoen AW, McCleland ML, O’Farrell PH (2010) Developmental control of late replication and S phase length. Curr Biol 20(23):2067–2077. doi:10.1016/j.cub.2010.10.021
Lis JT (2007) Imaging Drosophila gene activation and polymerase pausing in vivo. Nature 450(7167):198–202. doi:10.1038/nature06324
Ebert A, Lein S, Schotta G, Reuter G (2006) Histone modification and the control of heterochromatic gene silencing in Drosophila. Chromosome Res 14(4):377–392. doi:10.1007/s10577-006-1066-1
Zielke N, Kim KJ, Tran V, Shibutani ST, Bravo MJ, Nagarajan S, van Straaten M, Woods B, von Dassow G, Rottig C, Lehner CF, Grewal SS, Duronio RJ, Edgar BA (2011) Control of Drosophila endocycles by E2F and CRL4(CDT2). Nature 480(7375):123–127. doi:10.1038/nature10579
Klapholz B, Dietrich BH, Schaffner C, Heredia F, Quivy JP, Almouzni G, Dostatni N (2009) CAF-1 is required for efficient replication of euchromatic DNA in Drosophila larval endocycling cells. Chromosoma 118(2):235–248. doi:10.1007/s00412-008-0192-2
Calado R, Young N (2012) Telomeres in disease. F1000 Med Rep 4:8. doi:10.3410/M4-8
Mozgova I, Mokros P, Fajkus J (2010) Dysfunction of chromatin assembly factor 1 induces shortening of telomeres and loss of 45S rDNA in Arabidopsis thaliana. Plant Cell 22(8):2768–2780. doi:10.1105/tpc.110.076182
Sedkov Y, Cho E, Petruk S, Cherbas L, Smith ST, Jones RS, Cherbas P, Canaani E, Jaynes JB, Mazo A (2003) Methylation at lysine 4 of histone H3 in ecdysone-dependent development of Drosophila. Nature 426(6962):78–83. doi:10.1038/nature02080
Linger J, Tyler JK (2005) The yeast histone chaperone chromatin assembly factor 1 protects against double-strand DNA-damaging agents. Genetics 171(4):1513–1522. doi:10.1534/genetics.105.043000
Kirik A, Pecinka A, Wendeler E, Reiss B (2006) The chromatin assembly factor subunit FASCIATA1 is involved in homologous recombination in plants. Plant Cell 18(10):2431–2442. doi:10.1105/tpc.106.045088
Endo M, Ishikawa Y, Osakabe K, Nakayama S, Kaya H, Araki T, Shibahara K, Abe K, Ichikawa H, Valentine L, Hohn B, Toki S (2006) Increased frequency of homologous recombination and T-DNA integration in Arabidopsis CAF-1 mutants. The EMBO journal 25(23):5579–5590. doi:10.1038/sj.emboj.7601434
Song Y, He F, Xie G, Guo X, Xu Y, Chen Y, Liang X, Stagljar I, Egli D, Ma J, Jiao R (2007) CAF-1 is essential for Drosophila development and involved in the maintenance of epigenetic memory. Dev Biol 311(1):213–222. doi:10.1016/j.ydbio.2007.08.039
Schuermann D, Molinier J, Fritsch O, Hohn B (2005) The dual nature of homologous recombination in plants. Trends Genet 21(3):172–181. doi:10.1016/j.tig.2005.01.002
Friesner J, Britt AB (2003) Ku80- and DNA ligase IV-deficient plants are sensitive to ionizing radiation and defective in T-DNA integration. Plant J 34(4):427–440
Downs JA, Kosmidou E, Morgan A, Jackson SP (2003) Suppression of homologous recombination by the Saccharomyces cerevisiae linker histone. Mol Cell 11(6):1685–1692
Prado F, Aguilera A (2005) Partial depletion of histone H4 increases homologous recombination-mediated genetic instability. Mol Cell Biol 25(4):1526–1536. doi:10.1128/MCB.25.4.1526-1536.2005
Fischer S, Prykhozhij S, Rau MJ, Neumann CJ (2007) Mutation of zebrafish caf-1b results in S phase arrest, defective differentiation, and p53-mediated apoptosis during organogenesis. Cell Cycle 6(23):2962–2969
Wen P, Quan Z, Xi R (2012) The biological function of the WD40 repeat-containing protein p55/Caf1 in Drosophila. Dev Dyn 241(3):455–464. doi:10.1002/dvdy.23730
Hoek M, Stillman B (2003) Chromatin assembly factor 1 is essential and couples chromatin assembly to DNA replication in vivo. Proc Natl Acad Sci USA 100(21):12183–12188. doi:10.1073/pnas.1635158100
Quivy JP, Grandi P, Almouzni G (2001) Dimerization of the largest subunit of chromatin assembly factor 1: importance in vitro and during Xenopus early development. EMBO J 20(8):2015–2027. doi:10.1093/emboj/20.8.2015
Kaya H, Shibahara KI, Taoka KI, Iwabuchi M, Stillman B, Araki T (2001) FASCIATA genes for chromatin assembly factor-1 in arabidopsis maintain the cellular organization of apical meristems. Cell 104(1):131–142
Kats ES, Albuquerque CP, Zhou H, Kolodner RD (2006) Checkpoint functions are required for normal S-phase progression in Saccharomyces cerevisiae RCAF- and CAF-I-defective mutants. Proc Natl Acad Sci USA 103(10):3710–3715. doi:10.1073/pnas.0511102103
Haber JE, Braberg H, Wu Q, Alexander R, Haase J, Ryan C, Lipkin-Moore Z, Franks-Skiba KE, Johnson T, Shales M, Lenstra TL, Holstege FC, Johnson JR, Bloom K, Krogan NJ (2013) Systematic triple-mutant analysis uncovers functional connectivity between pathways involved in chromosome regulation. Cell reports 3(6):2168–2178. doi:10.1016/j.celrep.2013.05.007
Pires-daSilva A, Sommer RJ (2003) The evolution of signalling pathways in animal development. Nat Rev Genet 4(1):39–49. doi:10.1038/nrg977
Schonrock N, Exner V, Probst A, Gruissem W, Hennig L (2006) Functional genomic analysis of CAF-1 mutants in Arabidopsis thaliana. J Biol Chem 281(14):9560–9568. doi:10.1074/jbc.M513426200
Autran D, Baroux C, Raissig MT, Lenormand T, Wittig M, Grob S, Steimer A, Barann M, Klostermeier UC, Leblanc O, Vielle-Calzada JP, Rosenstiel P, Grimanelli D, Grossniklaus U (2011) Maternal epigenetic pathways control parental contributions to Arabidopsis early embryogenesis. Cell 145(5):707–719. doi:10.1016/j.cell.2011.04.014
Yatim A, Benne C, Sobhian B, Laurent-Chabalier S, Deas O, Judde JG, Lelievre JD, Levy Y, Benkirane M (2012) NOTCH1 nuclear interactome reveals key regulators of its transcriptional activity and oncogenic function. Mol Cell 48(3):445–458. doi:10.1016/j.molcel.2012.08.022
Mourikis P, Lake RJ, Firnhaber CB, DeDecker BS (2010) Modifiers of notch transcriptional activity identified by genome-wide RNAi. BMC Dev Biol 10:107. doi:10.1186/1471-213X-10-107
Mueller GC, Kajiwara K, Kim UH, Graham J (1978) Proposed coupling of chromatin replication, hormone action, and cell differentiation. Cancer Res 38(11 Pt 2):4041–4045
Tsubouchi T, Soza-Ried J, Brown K, Piccolo FM, Cantone I, Landeira D, Bagci H, Hochegger H, Merkenschlager M, Fisher AG (2013) DNA synthesis is required for reprogramming mediated by stem cell fusion. Cell 152(4):873–883. doi:10.1016/j.cell.2013.01.012
Chia G, Egli D (2013) Connecting the cell cycle with cellular identity. Cell Reprogram 15(5):356–366. doi:10.1089/cell.2013.0041
Newmark PA, Sanchez Alvarado A (2002) Not your father’s planarian: a classic model enters the era of functional genomics. Nat Rev Genet 3(3):210–219. doi:10.1038/nrg759
Wagner DE, Wang IE, Reddien PW (2011) Clonogenic neoblasts are pluripotent adult stem cells that underlie planarian regeneration. Science 332(6031):811–816. doi:10.1126/science.1203983
Newmark PA, Sanchez Alvarado A (2000) Bromodeoxyuridine specifically labels the regenerative stem cells of planarians. Dev Biol 220(2):142–153. doi:10.1006/dbio.2000.9645
Wenemoser D, Lapan SW, Wilkinson AW, Bell GW, Reddien PW (2012) A molecular wound response program associated with regeneration initiation in planarians. Genes Dev 26(9):988–1002. doi:10.1101/gad.187377.112
Kaufman PD, Kobayashi R, Kessler N, Stillman B (1995) The p150 and p60 subunits of chromatin assembly factor I: a molecular link between newly synthesized histones and DNA replication. Cell 81(7):1105–1114
Smith S, Stillman B (1991) Stepwise assembly of chromatin during DNA replication in vitro. EMBO J 10(4):971–980
Tyler JK, Collins KA, Prasad-Sinha J, Amiott E, Bulger M, Harte PJ, Kobayashi R, Kadonaga JT (2001) Interaction between the Drosophila CAF-1 and ASF1 chromatin assembly factors. Mol Cell Biol 21(19):6574–6584
Wei C, Liu J, Yu Z, Zhang B, Gao G, Jiao R (2013) TALEN or Cas9, rapid, efficient and specific choices for genome modifications. J Genet Genom 40(6):281–289. doi:10.1016/j.jgg.2013.03.013
Acknowledgments
This work was supported by grants from the National Natural Science Foundation of China (NSFC) [Nos. 31271573, 31228015, 31201007, 81470846] and the 973 Program [2012CB825504]. We are grateful to Gabriel Calvin and Jen Kennedy for critical reading of the manuscript. We thank the anonymous reviewers for constructive suggestions.
Author information
Authors and Affiliations
Corresponding author
Additional information
Z. Yu and J. Liu contributed equally to this work.
Rights and permissions
About this article
Cite this article
Yu, Z., Liu, J., Deng, WM. et al. Histone chaperone CAF-1: essential roles in multi-cellular organism development. Cell. Mol. Life Sci. 72, 327–337 (2015). https://doi.org/10.1007/s00018-014-1748-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-014-1748-3