Abstract
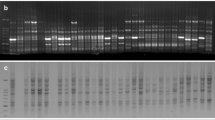
Hordeum vulgare (barley) and Triticum tauschii are related, but sexually incompatible, species. This study was conducted to determine the extent of homology between the genomes of barley and T. tauschii using a common set of restriction fragment length polymorphism (RFLP) markers. Results showed that >95% of low-copy sequences are shared, but 42% of the conserved sequences showed copy-number differences. Sixty-three loci were mapped in T. tauschii using RFLP markers previously mapped in barley. A comparison of RFLP marker order showed that, in general, barley and T. tauschii have conserved linkage groups, with markers in the same linear orders. However, six of the seven linkage groups of T. tauschii contained markers which mapped to unrelated (i.e., non-homoeologous) barley chromosomes. Additionally, four of the T. tauschii linkage groups contained markers that were switched in order with respect to barley. All the chromosome segments differing between T. tauschii and barley contained markers that were detected by multi-copy probes. The results suggest that the observed differences between the T. tauschii and barley genomes were brought about by duplications or deletions of segments in one or both species. The implications of these findings for genetic mapping, breeding, and plant genome evolution are discussed.
Similar content being viewed by others
References
Ahn S, Tanksley SD (1993) Comparative linkage maps of the rice and maize genomes. Proc Natl Acad Sci USA 90:7980–7984
Ahn S, Anderson JA, Sorrells ME, Tanksley SD (1993) Homoeologous relationships of rice, wheat and maize chromosomes. Mol Gen Genet 241:483
Arnheim N (1983) Concerted evolution of multigene families. In: Koehn R, Nei M (eds) Evolution of genes and proteins. Sinauer, Sunderland, pp. 38–61
Beltz G, Jacobs K, Eickbush T, Cherbas P, Kafatos F, (1983) Isolation of multigene families and determination of homologies by filter hybridization methods. Methods Enzymol 100:266–285
Benito C, Figueiras AM, Gonzalez-Jaen MT, Salinas J (1985) Biochemical evidence of homoeology between wheat and barley chromosomes. Z. Pflanzenzuchtg 94:307–320
Bennett MD, Smith JB (1976) Nuclear DNA amounts in angiosperms. Philos Trans R Soc Lon (Biol) 274:227–274
Bernatzky R, Tanksley SD (1986) Methods for detection of singleor low-copy sequences in tomato on Southern blots. Plant Mol Biol Rep 4:37–41
Bonierbale MW, Plaisted RL, Tanksley SD (1988) RFLP maps based on a common set of clones reveal modes of chromosomal evolution in potato and tomato. Genetics 120:1095–1103
Chao S, Sharp PJ, Worland AJ, Warham EJ, Koebner RMD, Gale MD (1989) RFLP-based genetic maps of wheat homoeologous group-7 chromosomes. Theor Appl Genet 78:495–504
Chen F, Hayes PM (1989) A comparison of Hordeum bulbosum-mediated haploid production efficiency in barley using in vitro floret and tiller culture. Theor Appl Genet 77:701–704
Devos KM, Atkinson MD, Chinoy CN, Francis HA, Harcourt RL, Koebner RMD, Liu CJ, Masojc P, Xie DX, Gale MD (1993) Chromosomal rearrangements in the rye genome relative to that of wheat. Theor Appl Genet 85:673–680
Dover G (1982) Molecular drive: a cohesive mode of species evolution. Nature 284:111–117
Feinberg AP, Vogelstein B (1983) A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity. Anal Biochem 132:6–13
Freeling M (1984) Plant transposable elements and insertion sequences. Annu Rev Plant Physiol 35:277–298
Gebhardt C, Ritter E, Barone A, Debener T, Walkemeier B, Schachtschabel U, Kaufmann, H, Thompson RD, Bonierbale MW, Ganal MW, Tanksley SD, Salamini F (1991) RFLP maps of potato and their alignment with the homoeologus tomato genome. Theor Appl Genet 83:49–57
Gill KS, Lubbers EL, Gill BS, Raupp WJ, Cox TS (1991) A genetic linkage map of Triticum tauschii (DD) and its relationship to the D genome of bread wheat (AABBDD). Genome 34:362–374
Harberd NP, Flavell RB, Thompson RD (1987) Identification of a transposon-like insertion in a Gen-1 allele of wheat. Mol Gen Genet 209:326–332
Hart GE (1987) Genetic and biochemical studies of enzymes, In: Heyne E. (ed) Wheat and wheat improvement. American Society of Agronomy, Madison, Wisconsin, pp 199–214
Helentjaris T, Weber D, Wright S (1988) Identification of the genomic locations of duplicate nucleotide sequences in maize by restriction length polymorphisms. Genetics 118:353–363
Heun M, Kennedy AE, Anderson JA, Lapitan NLV, Sorrells ME, Tanksley SD (1991) Construction of an RFLP map for barley (Hordeum vulgare). Genome 34:437–447
Hulbert SH, Richter TE, Axtell JD, Bennetzen JL (1990) Genetic mapping and characterization of sorghum and related crops by means of maize DNA probes. Proc Natl Acad Sci USA 87:4251–4255
Kam-Morgan LNW, Gill BS, Muthukrishanan S (1989) DNA restriction fragment length polymorphisms: a strategy for genetic mapping of D genome of wheat. Genome 32:724–732
Kihara H (1944) Discovery of the DD-analyser, one of the ancestors of vulgare wheats. Agric Hort (Tokyo) 19:889–890
Kleinhofs A, Kilian A, Saghai Maroof MA, Biyashev RM, Hayes P, Chen FQ, Lapitan N, Fenwick A, Blake T, Kanazin V, Ananiev E, Dahleen L, Kudrna D, Bollinger J, Knapp SJ, Liu B, Sorrells M, Heun M, Franckowiak JD, Hoffman D, Skadsen R, Steffenson BJ (1993) A molecular, isozyme and morphological map of the barley (Hordeum vulgare) genome. Theor Appl Genet 86:705–712
Kurata N, Moore G, Foote T, Yano M, Minobe Y, Gale M (1994) Conservation of genome structure between rice and wheat. Biol Technology 12:276–278
Lander ES, Green P, Abrahamson J, Barlow A, Daly MJ, Lincoln SE, Newburg L (1987) MAPMAKER: An interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Liao YC, Niks RE (1991) Application of a set of 14 cDNA probes from wheat to detect restriction fragment length polymorphism (RFLP) in barley. Euphytica 53:115–119
Liu CJ, Atkinson MD, Chinoy CN, Devos KM, Gale MD (1992) Nonhomoeologous translocations between group 4, 5, and 7 chromosomes within wheat and rye. Theor Appl Genet 83:305–312
Liu YC, Ikeda TM, Tsunewaki K (1992) Moderately repeated, dispersed, and highly variable (MRDHV) genomic sequences of common wheat usable for cultivar identification. Theor Appl Genet 84:535–543
McCouch SR, Kochert G, Yu ZH, Wang ZY, Khush GS, Coffman WR, Tanksley SD (1988) Molecular mapping of rice chromosomes. Theor Appl Genet 76:815–829
McFadden ES, Sears ER (1944) The artificial synthesis of Triticum Spelta. Rec Genet Soc Am 13:26–27
McFadden ES, Sears ER (1946) The origin of Triticum spelta and its free-threshing hexaploid relatives. J Hered 37:81–89, 107–116
Nkongolo KK, Lapitan NLV, Quick JS, Muhlmann MD (1993) An optimized fluorescence in situ hybridization procedure for detecting rye chromosomes in wheat. Genome 36:701–705
Sharp PJ, Chao S, Desao S, Gale MD (1989) The isolation, characterization and application in the Triticeae of a set of wheat RFLP probes identifying each homoeologous chromosome arm. Theor Appl Genet 78:342–348
Smith G (1975) Evolution of repeated DNA sequence by unequal crossover. Science 191:528–535
Tanksley SD, Bernatzky R, Lapitan NL, Prince JP (1988) Conservation of gene repertoire but not gene order in pepper and tomato. Proc Natl Acad Sci USA 85:6419–6423
Author information
Authors and Affiliations
Additional information
Communicated by K. Tsunewaki
Published with the approval of the Director of the Colorado State University/Agricultural Experiment Station
Rights and permissions
About this article
Cite this article
Namuth, D.M., Lapitan, N.L.V., Gill, K.S. et al. Comparative RFLP mapping of Hordeum vulgare and Triticum tauschii . Theoret. Appl. Genetics 89, 865–872 (1994). https://doi.org/10.1007/BF00224511
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00224511