Abstract
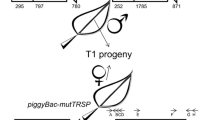
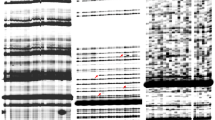
In Antirrhinum majus only autonomous Tam3 transposons have been characterized. We investigated whether an artificial dTam3 element, with a deletion in the presumptive transposase coding region, can be trans-activated in tobacco by an activator Tam3 element, which was immobilized by the deletion of one inverted repeat. A phenotypic assay based on restored hygromycin resistance demonstrates that a dTam3 element harbouring a bacterial plasmid can be trans-activated with a low frequency. Molecular analysis confirms that the dTam3 element has been excised from the HPTII marker gene. Reintegration of the dTam3 element into the tobacco genome is detected only in one out of six hygromycin-resistant plants analysed. PCR analysis of empty donor sites shows that excision of the dTam3 element in tobacco results in rearrangements (deletions and additions), that have been shown to be characteristic of Tam3 excision in the original host Antirrhinum majus. This trans-activation assay allowed us to establish that, in contrast to what has been detected in Antirrhinum majus, a periodical temperature shift down to 15°C does not enhance dTam3 transposition in regenerating tobacco calli.
Similar content being viewed by others
References
BakerB, CouplandG, FedoroffN, StarlingerP, SchellJ: Phenotypic assay for excision of the maize controlling element Ac in tobacco. EMBO J 6: 1547–1554 (1987).
CarpenterR, MartinC, CoenES: Comparison of genetic behaviour of the transposable element Tam3 at two unlinked pigment loci in Antirrhinum majus. Mol Gen Genet 207: 82–89 (1987).
Coen ES, Robbins TP, Almeida J, Hudson A, Carpenter R: Consequences and mechanisms of transposition in Antirrhinum majus. In: Berg DE, Howe MM (eds) Mobile DNA, pp 413–436. American Society for Microbiology (1989).
CouplandG, BakerB, SchellJ, StarlingerP: Characterization of the maize transposable element Ac by internal deletions. EMBO J 7: 3653–3659 (1988).
FedoroffNV, FurtekDB, NelsonOEJr: Cloning of the bronze locus in maize by a simple and generalizable procedure using the transposable controlling element Activator (Ac). Proc Natl Acad Sci USA 81: 3825–3829 (1984).
HaringMA, GaoJ, VolbedaTj, RommensCMT, NijkampHJJ, HilleJ: A comparative study of Tam3 and Ac transposition in transgenic tobacco and petunia plants. Plant Mol Biol 13: 189–201 (1989).
HehlR, BakerB: Induced transposition of Ds by a stable Ac in crosses of transgenic tobacco plants. Mol Gen Genet 217: 53–59 (1989).
JonesJDG, CarlandFM, MaligaP, DoonerHK: Visual detection of transposition of the maize element Activator in tobacco seedlings. Science 244: 204–207 (1989).
KnappS, CouplandG, UhrigH, StarlingerP, SalaminiF: Transposition of the maize transposable element Ac in Solanum tuberosum. Mol Gen Genet 213: 285–290 (1988).
LassnerMW, PalysM, YoderJI: Genetic transactivation of Dissociation elements in transgenic tomato plants. Mol Gen Genet 218: 25–32 (1989).
ListerC, MartinC: Molecular analysis of a transposon-induced deletion of the nivea locus in Antirrhinum majus. Genetics 123: 417–425 (1989).
MartinC, CarpenterR, SommerH, SaedlerH, CoenEC: Molecular analysis of instability in flower pigmentation of Antirrhinum majus, following isolation of the pallida locus by transposon tagging. EMBO J 4: 1625–1630 (1985).
MartinC, PrescottA, ListerC, MackayS: Activity of the transposon Tam3 in Antirrhinum majus and tobacco: possible role of DNA methylation. EMBO J 8: 997–1004 (1989).
MartinCR, MackayS, CarpenterR: Large scale chromosomal restructuring is induced by the transposable element Tam3 at the nivea locus of Antirrhinum majus. Genetics 119: 171–184 (1988).
MastersonRV, FurtekDB, GreveldingC, SchellJ: A maize Ds transposable element containing a dihydrofolate reductase gene transposes in Nicotiana tabacum and Arabidopsis thaliana. Mol Gen Genet 219: 461–466 (1989).
MottoM, MaddaloniM, PonzianaG, BrembillaM, MarottaR, DiFonzoN, SoaveC, ThompsonR, SalaminiF: Molecular cloning of the o2-m5 allele of Zea mays using transposon marking. Mol Gen Genet 212: 488–494 (1988).
NeversP, ShephardN, SaedlerH: Plant transposable elements. Adv Bot Res 12: 103–203 (1986).
RobbinsTP, CarpenterR, CoenES: A chromosome rearrangement suggests that donor and recipient sites are associated during Tam3 transposition in Antirrhinum majus. EMBO J 8: 5–13 (1989).
SaikiRK, GelfandDH, StoffelS, ScharfSJ, HiguchiR, HornGT, MullisKB, ErlichHA: Primer directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science 239: 487–491 (1988).
SommerH, CarpenterR, HarrisonBJ, SaedlerH: The transposable element Tam3 of Antirrhinum majus generates a novel type of sequence alterations upon excision. Mol Gen Genet 199: 225–231 (1985).
SpenaA, AalenRB, SchulzeSC: Cell-autonomous behaviour of the rolC gene of Agrobacterium rhizogenes during leaf development: A visual assay for transposition excision in transgenic plants. Plant Cell 1: 1157–1164 (1989).
VanSluysMA, TempéJ, FedoroffN: Studies on the introduction and mobility of the maize Activator element in Arabidopsis thaliana and Daucus carota. EMBO J 6: 3881–3889 (1987).
YoderJI, PalysJ, AlpertK, LassnerM: Ac transposition in transgenic tomato plants. Mol Gen Genet 213: 291–296 (1988).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Haring, M.A., Teeuwen-de Vroomen, M.J., Nijkamp, H.J.J. et al. Trans-activation of an artificial dTam3 transposable element in transgenic tobacco plants. Plant Mol Biol 16, 39–47 (1991). https://doi.org/10.1007/BF00017915
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00017915