Abstract
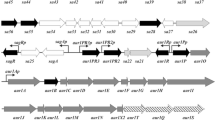
AStreptomyces aureofaciens gene,gap, encoding glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was previously identified. Hybridization studies suggested the presence of a secondgap gene inS. aureofaciens. To clone the gene,S. aureofaciens subgenomic library was screened with an oligonucleotide probe encoding a peptide motif conserved in all GAPDH. 3352 bp positiveBamHI fragment was identified, the length of which correlated with the hybridization signal. The nucleotide sequence of the fragment was determined, and analysis of the sequence revealed the presence of three open reading frames (ORF). However, none of the genes coded for GAPDH. All three genes formed an operon, consisting of geneorf251, with a high homology to a conserved gene present only in archæabacteria, and thealdA andadhA genes homologous to various eukaryotic and prokaryotic aldehyde- and alcohol-dehydrogenases, with maximum homology to the phenylacetaldehyde dehydrogenases and arylalcohol dehydrogenases, respectively.
Similar content being viewed by others
References
Altschul S.F., Gish W., Miller W., Myers E.w., Lipman D.J.: Basic local alignment search tool.J. Mol. Biol. 215, 403–410 (1990).
Ausubel F.M., Brent R., Kingston R.E., Moore D.O., Seidman J.S., Smith J.A. Struhl K.:Current Protcols in Molecular Biology. Wiley, New York 1987.
Beltrametti F., Marconi A.M., Bestetti G., Colombo C., Galli E., Ruzzi M., Zennaro E.: Sequencing and functional analysis of styrene catabolism genes fromPseudomonas fluorescens ST.Appl. Environ. Microbiol. 63, 2232–2239 (1997).
Bibb M.J., Cohen S.N.: Gene expression inStreptomyces: construction and application of promoter-probe plasmid vectors inStreptomyces lividans.Mol. Gen. Genet. 187, 265–277 (1982).
Bibb M.J., Findlay P.R., Johnson M.W.: The relationship between base composition and codon usage in bacterial genes and its use for the simple and reliable identification of protein-coding sequences.Gene 30, 157–166 (1984).
Brunner N.A., Brinkmann H., Siebers B., Hensel R.: NAD+-dependent glyceraldehyde-3-phosphate dehydrogenase fromThermoproteus tenax.J. Biol. Chem. 273, 6149–6156 (1998).
Chater K.F., Hopwood D.A.: Streptomyces, pp. 83–99 in A.L. Sonenshein, J.A. Hoch, R. Losick (Eds):Bacillus subtilis and Other Gram-Positive Bacteria: Biochemistry, Physiology and Molecular Genetics. American Society for Microbiology, Washington (DC) 1993.
Eaton R.W.:p-Cumate catabolic pathway inPseudomonas putida F1: cloning and characterization of DNA carrying thecmt operon.J. Bacteriol. 178, 1351–1362 (1996).
Ferrandez A., Prieto M.A., Garcia J.L., Diaz E.: Molecular characterization of PadA, a phenylacetaldehyde dehydrogenase fromEscherichia coli.FEBS Lett. 7, 23–27 (1997).
Freiberg C., Fellay R., Bairoch A., Broughton W.J., Rosenthal A., Perret X.: Molecular basis of symbiosis betweenRhizobium and legumes.Nature 387, 394–401 (1997).
Gillooly D.J., Robertson A.G., Fewson C.A.: Molecular characterization of benzyl alcohol dehydrogenase and benzaldehyde dehydrogenase II ofAcinetobacter calcoaceticus.Biochem. J. 330, 1375–1381 (1998).
Habenicht A.: The non-phosphorylating glyceraldehyde-3-phosphate dehydrogenase: biochemistry, structure, occurrence and evolution.Biol. Chem. 378, 1413–1419 (1997).
Harayama S., Rekik M., Wubbolts M., Rose K., Leppik R.A., Timmis K.N.: Characterization of five genes in the upper-pathway operon of TOL plasmid pWW0 fromPseudomonas putida and identification of the gene productsJ. Bacteriol. 171, 5048–5055 (1989).
Hempel J., Nicholas H., Lindahl R.: Aldehyde dehydrogenases: widespread structural and functional diversity within a shared framework.Prot. Sci. 2, 1890–1900 (1993).
Hidalgo E., Chen Y.M., Lin E.C., Aguilar J.: Molecular cloning and DNA sequencing of theEscherichia coli K-12ald gene encoding aldehyde dehydrogenase.J. Bacteriol. 173, 6118–6123 (1991).
Hopwood D.A., Bibb M.J., Chater K.F., Kieser T., Bruton C.J., Kieser H.M., Lydiate D.L., Smith C.P., Ward J.M., Schrempf H.:Genetic Manipulation of Streptomyces. A Laboratory Manual. The John Innes Foundation, Norwich 1985.
Hurley T.D., Bosron W.F., Hamilton J.A., Amzel L.M.: Structure of human b1 b1 alcohol dehydrogenase: catalytic effects of non-active-site substitutions.Proc. Nat. Acad. Sci. USA 88, 8149–8153 (1991).
James K.D., Williams P.A.:ntn genes determining the early steps in the divergent catabolism of 4-nitrotoluen and toluen inPseudomonas sp. strain TW3.J. Bacteriol. 180, 2043–2049 (1998).
Junker F., Kiewitz R., Cook A.M.: Characterization of thep-toluenesulfonate operontsaMBCD andtsaR inComamonas testosteroni T-2.J. Bacteriol. 179, 919–927 (1997).
Jornvall H., Persson B., Jeffrey J.: Characteristics of alcohol/polyol dehydrogenases. The of zinc-containing long-chain alcohol dehydrogenases.Eur. J. Biochem. 167, 195–201 (1987).
Kormanec J., Řežuchová B., Farkašovský M.: Optimization ofStreptomyces aureofaciens transformation and disruption of thehrdA gene encoding a homologue of the principal s factor.J. Gen. Microbiol. 139, 2525–2529 (1993).
Kormanec J., Lempelová A., Farkašovský M., Homérová D.: Cloning, sequencing and expression inEscherichia coli of aStreptomyces aureofaciens gene encoding glyceraldehyde-3-phosphate dehydrogenase.Gene 165, 77–80 (1995).
Kormanec J., Ševčíková B., Sprušanský O, Benada O., Kofroňová O., Nováková R., Řežuchová B., Potúčková L., Homérová D.: TheStreptomyces aureofaciens homologue of thewhiB gene is essential for sporulation; its expression c correlates with the developmental stage.Folia Microbiol. 43, 605–612 (1998).
Liu Z.-J., Sun Y.-J., Rose J., Chung Y.-J., Hsiao C.-D., Chang W.-R., Kuo I., Perozich J., Lindahl R., Hempel J., Wang B.-C.: The first structure of an aldehyde dehydrogenase reveals novel interactions between NAD and the Rossemann fold.Nat. Struct. Biol. 4, 317–326 (1997).
Maxam A.M., Gilbert W.: Sequencing end-labelled DNA with base specific chemical cleavages.Methods Enzymol. 65, 449–560 (1980).
Shaw J.P., Rekik M., Schwager F., Harayama S.: Kinetic studies on benzyl alcohol dehydrogenase encoded by TOL plasmid pWWO. A member of the zinc-containing long chain alcohol dehydrogenase family.J. Biol. Chem. 268, 10842–10850 (1993).
Skarzyński T., Moody P.C.E., Wonacott A.J.: Structure of holo-glyceraldehyde-3-phosphate dehydrogenase fromBacillus stearothermophilus at 1.8 Å resolution.J. Mol. Biol. 193, 171–187 (1987).
Steinmetz C.G., Xie P., Weiner H., Hurley T.D.: Structure of mitochondrial aldehyde dehydrogenase: the genetic component of ethanol aversion.Structure 5, 701–711 (1997).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sprušanský, O., Homérová, D., Ševčíková, B. et al. Cloning of the putative aldehyde dehydrogenase,aldA, gene fromStreptomyces aureofaciens . Folia Microbiol 44, 491–502 (1999). https://doi.org/10.1007/BF02816249
Received:
Revised:
Issue Date:
DOI: https://doi.org/10.1007/BF02816249