Abstract
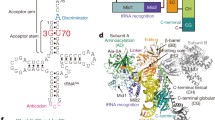
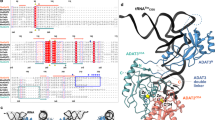
It has previously been shown that the single mutation E222K in glutaminyl-tRNA synthetase (GlnRS) confers a temperature-sensitive phenotype onEscherichia coli. Here we report the isolation of a pseudorevertant of this mutation, E222K/C171G, which was subsequently employed to investigate the role of these residues in substrate discrimination. The three-dimensional structure of the tRNAGln: GlnRS:ATP ternary complex revealed that both E222 and C171 are close to regions of the protein involved in interactions with both the acceptor stem and the 3′ end of tRNAGln. The potential involvement of E222 and C171 in these interactions was confirmed by the observation that GlnRS-E222K was able to mischargesupF tRNATyr considerably more efficiently than the wild-type enzyme, whereas GlnRS-E222K/C171G could not. These differences in substrate specificity also extended to anticodon recognition, with the double mutant able to distinguishsupE tRNA GlnCUA from tRNA Gln2 considerably more efficiently than GlnRS E222K. Furthermore, GlnRS-E222K was found to have a 15-fold higher Km for glutamine than the wild-type enzyme, whereas the double mutant only showed a 7-fold increase. These results indicate that the C171G mutation improves both substrate discrimination and recognition at three domains in GlnRS-E222K, confirming recent proposals that there are extensive interactions between the active site and regions of the enzyme involved in tRNA binding.
Similar content being viewed by others
References
Bhattacharyya T, Roy S (1993) A fluorescence spectroscopic study of substrate-induced conformational changes in glutaminyl-tRNA synthetase. Biochemistry 32:9268–9273
Englisch-Peters S, Conley J, Plumbridge J, Leptak C, Söll D, Rogers MJ (1991) Mutant enzymes and tRNAs as probes of the glutaminyl-tRNA synthetase: tRNAGln interaction. Biochimie 73:1501–1508
Eriani G, Delarue M, Poch O, Gangloff J, Moras D (1990) Partition of tRNA synthetases into two classes based on mutually exclusive sets of sequence motifs. Nature 347:203–206
Hayase Y, Jahn M, Rogers MJ, Sylvers LA, Koizumi M, Inoue H, Ohtsuka E, Söll D (1992) Recognition of bases inEscherichia coli tRNAGln by glutaminyl-tRNA synthetase: a complete identity set. EMBO J 11:4159–4165
Hoben P, Söll D (1985) Glutaminyl-tRNA synthetase ofEscherichia coli. Methods Enzymol 113:55–59
Hoben P, Royal N, Cheung A, Yamao F, Biemann K, Söll D (1982)Escherichia coli glutaminyl-tRNA synthetase. II. Characterization of theglnS gene product. J Biol Chem 257:11644–11650
Hong KW, Ibba M, Weygand-Durasevic I, Rogers MJ, Thomann HU, Söll D (1996) Transfer RNA-dependent cognate amino acid recognition by an aminoacyl-tRNA synthetase. EMBO J 15:1983–1991
Inokuchi H, Celis JE, Smith JD (1974) Mutant tyrosine transfer ribonucleic acids ofEscherichia coli: construction by recombination of a double mutant AlG82 chargeable with glutamine. J Mol Biol 85:187–192
Inokuchi H, Kodaira M, Yamao F, Ozeki H (1979) Identification of transfer RNA suppressors inEscherichia coli. II. Duplicate genes for tRNA Gln2 . J Mol Biol 132:663–677
Inokuchi H, Hoben P, Yamao F, Ozeki H, Söll D (1984) Transfer RNA mischarging mediated by a mutantEscherichia coli glutaminyl-tRNA synthetase. Proc Natl Acad Sci USA 81:5076–5080
Jahn M, Rogers MJ, Söll D (1991) Anticodon and acceptor stem nucleotides in tRNAGln are major recognition elements forE. coli glutaminyl-tRNA synthetase. Nature 352:258–269
Miller JH (1992) A short course in bacterial genetics. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Mullis KB, Ferre F, Gibbs RA (1994) The polymerase chain reaction. Birkhauser press, Boston
Perona JJ, Swanson RN, Rould MA, Steitz TA, Söll D (1989) Structural basis for misaminoacylation by mutantE. coli glutaminyl-tRNA synthetase enzymes. Science 246:1152–1154
Rogers MJ, Weygand-Durasevic I, Schwob E, Sherman JM, Rogers KC, Adachi T, Inokuchi H, Söll D (1993) Selectivity and specificity in the recognition of tRNA byE. coli glutaminyl-tRNA synthetase. Biochimie 75:1083
Rogers MJ, Adachi T, Inokuchi H, Söll D (1994) Functional communication in the recognition of tRNA byEscherichia coli glutaminyl-tRNA synthetase. Proc Natl Acad Sci USA 91:291–295
Rould MA, Perona JJ, Söll D, Steitz TA (1989) Structure ofE. coli glutaminyl-tRNA synthetase complexed with tRNAGln and ATP at 2.8 Å resolution. Science 246:1135–1142
Rould MA, Perona JJ, Steitz TA (1991) Structural basis of anticodon loop recognition by glutaminyl-tRNA synthetase. Nature 352:213–218
Schulman LH (1991) Recognition of tRNAs by aminoacyl-tRNA synthetases. Prog Nucleic Acid Res Mol Biol 41:23–87
Smith JD, Celis JE (1973) Mutant tyrosine transfer RNA that can be charged with glutamine. Nature New Biol 243:66–71
Thomann HU, Ibba M, Hong KW, Söll D (1996) Homologous expression and purification of mutants of an essential protein by reverse epitope-tagging. Biotechnology 14:50–55
Yamao F, Inokuchi H, Cheung A, Ozeki H, Söll D (1982)Escherichia coli glutaminyl-tRNA synthetase. I. Isolation and DNA sequence of theglnS gene. J Biol Chem 257:11639–11643
Young RA, Davis RW (1983) Efficient isolation of genes by using antibody probes. Proc Natl Acad Sci USA 80:1194
Author information
Authors and Affiliations
Additional information
Communicated by K. Isono
Rights and permissions
About this article
Cite this article
Kitabatake, M., Inokuchi, H., Ibba, M. et al. Genetic analysis of functional connectivity between substrate recognition domains ofEscherichia coli glutaminyl-tRNA synthetase. Molec. Gen. Genet. 252, 717–722 (1996). https://doi.org/10.1007/BF02173978
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF02173978