Summary
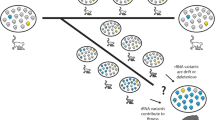
DNA restriction endonuclease fragment analysis is used to examine the genetic organization, inheritance and linkage associations of the ribosomal DNA in pea. The substantial variation observed in the length of the intergenic spacer region is shown to segregate in Mendelian fashion involving two independent genetic loci, designated Rrn1 and Rrn2. Linkage between Rrn1 and two marker loci on chromosome 4 establishes the approximate location of this tandem array. Rrn2 shows linkage with a set of isozyme loci which assort independently of other markers on all seven chromosomes. Combining these observations with previous cytological data, we suggest that Rrn2 and the isozyme loci linked to it constitute a new linkage group on chromosome 7. The general absence of spacer length classes common to both rRNA loci in any of the lines we examined indicates that little or no genetic exchange occurs between the nonhomologous nucleolar organizer regions.
Similar content being viewed by others
References
Appels R, Dvorak J (1982) The wheat ribosomal DNA spacer region: its structure and variation in populations and among species. Theor Appl Genet 63:337–348
Appels R, Gerlach WL, Dennis ES, Swift H, Peacock WJ (1980) Molecular and chromosomal organization of DNA sequences coding for the ribosomal RNAs in cereals. Chromosoma 78:293–311
Arnheim N, Krystal M, Schmickel R, Wilson G, Ryder O, Zimmer E (1980) Molecular evidence for genetic exchanges among ribosomal genes on nonhomologous chromosomes in man and apes. Proc Natl Acad Sci USA 77:7323–7327
Arnheim N, Treco D, Taylor B, Eicher EM (1982) Distribution of ribosomal gene length variants among mouse chromosomes. Proc Natl Acad Sci USA 79:4677–4680
Blixt S (1974) The pea. In: King RC (ed) Handbook of genetics, vol 2. Plenum, New York, pp 181–221
Boncinelli E, Borghese A, Graziani F, La Mantia G, Manzi A, Mariani C, Simeone A (1983) Inheritance of the rDNA spacer in D. melanogaster. Mol Gen Genet 189:370–374
Coen ES, Dover G (1983) Unequal exchanges and the coevolution of X and Y rDNA arrays in Drosophila melanogaster. Cell 33:849–855
Coen ES, Thoday JM, Dover G (1982) Rate of turnover of structural variants in the rDNA gene family of Drosophila melanogaster. Nature 295:564–568
Cuellar RE (1982) The structure and evolution of repeated DNA sequences in Pisum sativum: a molecular characterization. PhD Thesis, Stanford University
Delseny M, Aspart L, Cooke R, Grellet F, Penon P (1979) Restriction analysis of radish nuclear genes coding for rRNA: evidence for heterogeneity. Biochem Biophys Res Commun 91:540–547
Dover G (1982) Molecular drive: a cohesive mode of species evolution. Nature 299:111–117
Ellis THN, Davies DR, Castleton JA, Bedford ID (1984) The organization and genetics of rDNA length variants in pea. Chromosoma 91:74–81
Folkeson D (1984) Free segregation between a (3S–7S) interchange and genes within linkage group VII in Pisum sativum. Hereditas 101:227–233
Gerlach WL, Bedbrook JR (1979) Cloning and characterization of ribosomal RNA genes from wheat and barley. Nucleic Acids Res 7:1869–1885
Ingle J, Timmis JN, Sinclair J (1975) The relationship between satellite deoxyribonucleic acid, ribosomal ribonucleic acid gene redundancy, and genome size in plants. Plant Physiol 55:496–501
Jorgensen RA, Cuellar RE, Thompson WF (1982) Modes and tempos in the evolution of nuclear-encoded ribosomal RNA genes in legumes. Carnegie Inst Washington Yearb 81:98–101
Krystal M, D'Eustachio P, Ruddle FH, Arnheim N (1981) Human nucleolus organizers on nonhomologous chromosomes can share the same ribosomal gene variants. Proc Natl Acad Sci USA 78:5744–5748
Lamm R (1951) Cytological studies in Pisum. Hereditas 37:356–372
Lamm R (1981) Giemsa C-banding and silver-staining for cytological studies in Pisum. Hereditas 94:45–52
Lamppa GK, Honda S, Bendich AJ (1984) The relationship between ribosomal repeat length and genome size in Vicia. Chromosoma 89:1–7
Long EO, Dawid IB (1980) Repeated genes in eukaryotes. Annu Rev Biochem 49:727–764
Polans NO, Weeden NF, Thompson WF (1985) Inheritance, organization, and mapping of rbcS and cab multigene families in pea. Proc Natl Acad Sci USA 82:5083–5087
Reeder RH, Brown DD, Wellauer PK, Dawid IB (1976) Patterns of ribosomal DNA spacer lengths are inherited. J Mol Biol 105:507–516
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard RW (1984) Ribosomal DNA spacer length polymorphism in barley: Mendelian inheritance, chromosomal location and population dynamics. Proc Natl Acad Sci USA 81:8014–8018
Snape JW, Flavell RB, O'Dell M, Hughes WG, Payne PI (1985) Intrachromosomal mapping of the nucleolar organizer region relative to three marker loci on chromosome 1B of wheat (Triticum aestivum). Theor Appl Genet 69:263–270
Suiter KA, Wendel JF, Case JS (1983) LINKAGE-1: a PASCAL computer program for the detection and analysis of genetic linkage. J Hered 74:203–204
Treco D, Brownell E, Arnheim N (1982) The ribosomal gene nontranscribed spacer. In: Busch H, Rothblum L (eds) The cell nucleus, vol XII. Academic Press, New York, pp 101–126
Weeden N (1985) An isozyme linkage map for Pisum sativum. In: Hebblethwaite P, Dawkins T, Heath M (eds) The pea crop — a basis for improvement. Butterworths, Guildford, pp 55–66
Yakura K, Kato A, Tanifuji S (1984) Length heterogeneity of the large spacer of Vicia faba rDNA is due to the differing number of a 325 bp repetitive sequence elements. Mol Gen Genet 193:400–405
Author information
Authors and Affiliations
Additional information
Communicated by A. L. Kahler
Rights and permissions
About this article
Cite this article
Polans, N.O., Weeden, N.F. & Thompson, W.F. Distribution, inheritance and linkage relationships of ribosomal DNA spacer length variants in pea. Theoret. Appl. Genetics 72, 289–295 (1986). https://doi.org/10.1007/BF00288563
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00288563