Abstract
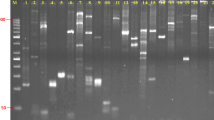
Genetic variability in six host genotype-specific pathotypes of pearl millet downy mildew pathogen S. graminicola was studied at the molecular level using mini- and micro-satellites. Our results indicated that microsatellites (GAA)6, (GACA)4, and especially (GATA)4 were quite informative and showed high levels of polymorphism among the pathotypes. The six pathotypes could be classified into five groups based on the cluster analysis of their genetic similarities, thereby confirming the existence of distinct host genotype-specific virulence in S. graminicola pathotypes. We demonstrate, for the first time, the use of DNA fingerprinting to detect genetic variation in downy mildew fungus of pearl millet.
Similar content being viewed by others
References
Anderson PA, Lawrence Pryor A (1992) Inheritance of restriction fragment length polymorphism in the flax rust Melamspora lini. Theor Appl Genet 84:845–850
Ball SL, Pike DJ (1984) Intercontinental variation in Sclerospora graminicola. Ann Appl Biol 104:41–51
Boeger JM, Chen RS, McDonald BA (1993) Gene flow between geographic populations of Mycosphaerella Graminicola (anamorph Septoria tritici) detected with restriction fragment length polymorphism markers. Phytopathology 83:1148–1154
Braithwaite KS, Manners JM (1989) Human hypervariable minisatellite probes detect DNA polymorphisms in the fungus Colletotrichum gloeosporioides. Curr Genet 16:473–475
Epplen JT (1988) On simple repeated GATA/GACA sequences in animal genomes: a critical reappraisal. J Hered 79:409–417
FAO (1983) 1982 FAO Production Year Book 36. FAO, Rome, Italy
Gupta VS, Ramakrishna W, Rawat SR, Ranjekar PK (1994) (CAC)5 detects DNA fingerprints and sequences homologous to gene transcripts in rice. Biochem Genet 32:1–8
Hammer JE, Farrall L, Orbach MJ, Valent B, Chumley FG (1989) Host species-specific conservation of a family of repeated DNA sequences in the genome of fungal plant pathogen. Proc Natl Acad Sci USA 86:9981–9985
Jeffreys AJ, Wilson V, Thein SL (1985) Individual specific “finger prints” of human DNA. Nature 316:76–79
Kashi Y, Tickochinsky Y, Genislav E, Traqi F, Nave A, Beckmann JS, Gruenbaum Y, Soller M (1990) Large restriction fragments containing poly TG are highly polymorphic in a variety of vertebrates. Nucleic Acids Res 18:1129–1132
Kohli Y, Morrall RA, Anderson JB, Kohn LM (1992) Local and trans-Canadian clonal distribution of Sclerotina sclerotiorum on canola. Phytopathology 82:875–880
Levy M, Rommao J, Marchetti MA, Hammer JE (1991) DNA fingerprinting with a dispersed repeated sequence resolves pathotype diversity in rice blast fungus. Plant Cell 3:95–102
Lieckfeldt E, Meyer W, Borner VT (1993) Rapid identification & differentiation of yeasts by DNA and PCR Fingerprinting. J Basic Microbiol 33:413–426
Longmire JL, Kraemer PM, Brown NC, Harde Kopf LC, Deaven LL (1990) A new multilocus DNA fingerprinting probe pV47–2. Nucleic Acids Res 18:1658
McDonald BA, Miles J, Nelson LR, Pettway RE (1994) Genetic variability in nuclear DNA in field populations of Stagonospora nodorum. Phytopathology 84:250–255
Meyer W, Koch A, Niemann C, Beyermann B, Epplen JT, Boerner T (1991) Differentiation of species and strains among filamentous fungi by DNA fingerprinting. Curr Genet 19:239–242
Meyer W, Mitchell TG, Freedman EZ, Vilgalys R (1993) Hybridization probes for conventional DNA Fingerprinting use as single primers in the polymerase chain reaction to distinguish strains of Cryptococcus neoformans. J Clin Microbiol 31: 2274–2280
Michelmore RN, Pawar MN, Williams RJ (1982) Heterothallism in Sclerospora graminicola. Phytopathology 72:1368–1372
Milgroom MG, Lipari SE, Powell WA (1992) DNA fingerprinting and analysis of population structure in the chestnut blight fungus Cryphonectria parasitica. Genetics 131:297–306
Rachie KO, Majumdar JV (1980) Pearl millet. Pennsylvania State University Press, Pa.
Ramakrishna W, Lagu MD, Gupta VS, Ranjekar PK (1994) DNA Fingerprinting rice in using oligonucleotide probes specific for simple repetitive DNA sequences. Theor Appl Genet 88: 402–406
Ramakrishna W, Chowdari KV, Lagu MD, Gupta VS, Ranjekar PK (1995) DNA fingerprinting to detect genetic variation in rice using hypervariable DNA sequences Theor Appl Genet 90:1000–1006
Sambrook J, Fritsch EF, Maniatis T (1989) Cloning: a laboratory manual 2nd edn. Cold Spring Harbor laboratory Press, Cold Spring Harbor, N. Y.
Schafer R, Zischler H, Birsner U, Becker A, Epplen JT (1988) Optimized oligonucleotide probes for DNA Fingerprinting. Electrophoresis 9:369–374
Singh SD, Singh G (1987) Resistance to downy mildew in pearl millet hybrid NHB 3. Indian Phytopathol 40:178–180
Singh SD, King SB, Werder J (1993) Downy mildew disease of pearl millet. Information bulletin No. 37. International Crops Research Institute for the Semi-Arid Tropics (ICRISAT), Patancheru A. P., India
Sivaramakrishnan S, Harikrishnan R, Thakur RP, Rao VP (1995) Use of RAPD markers for the detection of genetic variability in pearl millet downy mildew pathogen. In: Sharma RP, Chopra VL, Swaminathan MS (eds.). Proc 2nd Asia-Pacific Conference Agric Biotechno. Madras. Oxford & IBH Publishing Company, New Delhi (in press)
Tautz D, Renz M (1984) Simple sequence are ubiquitous repetitive components of eukaryotic genomes. Nucleic Acids Res 12: 4127–4138
Thakur RP (1993) Cereals Program, Annual Report 1992. Cereals Program, International Crops Research Institute for the Semi-Arid Tropics, Patancheru, A. P., India
Thakur RP, Shetty KG (1993) Variation in pathogenicity among single oospore isolates of S. graminicola, the causal organism of downy mildew in pearl millet. Plant Pathol 42:715–721
Thakur RP, Shetty KG, King SB (1992) Selection for host-specific virulence in asexual population of Sclerospora graminicola. Plant Path 41:626–632
Turner BC, Elder JF, Laughlin TF, Davis WP, Taylor DS (1992) Extreme clonal diversity & divergence in populations of a selfing hermaphroditic fish. Proc Natl Acad Sci USA 89: 10643–10647
Vassart G, Georges M, Monsieur R, Brocas H, Lequarre AS, Christopher D (1987) A sequence in M13 phage detects hypervariable minisatellites in humans & animal DNA. Science 235: 683–684
Vosman B, Avens P, Rus-Kortekaas W, Smulders MJM (1992) Identification of highly polymorphic DNA regions in tomato. Theor Appl Genet 85:239–244
Weising K, Ramser J, Kaemmer D, Kahl G, Epplen JT (1991a) Oligonucleotide fingerprinting in plants & fungi. In: Burke T, Dolf G, Jeffreys AJ, Wolff R (eds) DNA fingerprinting: approaches & application. Birkhauser, Basel, pp 312–319
Weising K, Kaemmer D, Epplen JT, Weigand F, Sexena M, Kahl G (1991b) DNA fingerprinting of Ascochyta rabiei with synthetic oligonucleotides. Curr Genet 19:483–489
Wetton JH, Carter RE, Parkin DT, Walters D (1987) Demographic study of a wild house sparrow population by DNA fingerprinting. Nature 327:147–149
Xia JQ, Correll JC, Lee FN, Marchetti MA, Rhoads DD (1993) DNA fingerprinting to examine the microgeographic variation in the Magnaporthe grisea (Pyricularia grisea) population in two rice fields in Arkansas. Phytopathology 83:1029–1035
Author information
Authors and Affiliations
Additional information
Communicated by J. Mac Key
Rights and permissions
About this article
Cite this article
Sastry, J.G., Ramakrishna, W., Sivaramakrishnan, S. et al. DNA fingerprinting detects genetic variability in the pearl millet downy mildew pathogen (Sclerospora graminicola). Theoret. Appl. Genetics 91, 856–861 (1995). https://doi.org/10.1007/BF00223892
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00223892