Abstract
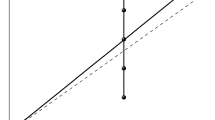
Through a simple computer simulation, we provide clear evidence that the base composition of nucleotide sequences should be taken into account to obtain accurate estimates of the genetic distances between homologous nucleotide sequences. We tested several stochastic models and demonstrated clearly that the inferred estimates of the number of substitutions are strongly dependent on the a priori conditions superimposed in the model, i.e., equiprobability in base composition and/or transition transversion ratio. In general, the more a priori assumptions there are, the more inaccurate the results. Whereas, rather accurate estimates are obtained with stochastic methods, which take into account base composition and do not superimpose unverified a priori conditions.
Similar content being viewed by others
References
Bernardi G (1993) The vertebrate genome: isochores and evolution. Mol Biol Evol 10(1):186–204
Bernardi, G., Olofsson B, Filipski J, Zerial M, Salinas J, Cuny G, Meunier-Rotival M, Rodier F (1985) The mosaic genome of warm-blooded vertebrates. Science 228:953–958
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Felsenstein J (1993) PHYLIP (phylogeny inference package). Department of Genetics, University of Washington, Seattle
Jukes, TH and Cantor CR (1969) Evolution of protein molecules. In: Munro HN (ed) Mammalian protein metabolism. Academic Press, New York, pp 21–132
Kimura, M. (1980) A simple method for estimating evolutionary rates of base substitution through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kishino, H., Hasegawa M (1989) Evaluation of the maximum likelihood estimate of the evolutionary tree topologies from DNA sequence data, and the branching order in Hominoidea. J Mol Evol 29:170–179
Lanave, C., Preparata G, Saccone C, Serio G (1984) A new method for calculating evolutionary substitution rates. J Mol Evol 20:86–93
Saccone C, Lanave C, Pesole G, Preparata G (1990) Influence of base composition on quantitative estimates of gene evolution. Methods Enzymol 183:570–583
Tajima F, Nei M (1984) Estimation of evolutionary distance between nucleotide sequences. Mol Biol Evol 1:269–285
Wolfe KH and Sharp PM (1993) Mammalian gene evolution: nucleotide sequence divergence between mouse and rat. J Mol Evol 37(4):441–456
Wolfe, KH, Sharp PM, Li W.-H (1989) Mutation rates differ among regions of the mammalian genome. Nature 337:283–285
Zharkikh A (1994) Estimation of evolutionary distances between nucleotide sequences. J Mol Evol 39(3):315–329
Author information
Authors and Affiliations
Additional information
Correspondence to: G. Pesole
Rights and permissions
About this article
Cite this article
Pesole, G., Dellisanti, G., Preparata, G. et al. The importance of base composition in the correct assessment of genetic distance. J Mol Evol 41, 1124–1127 (1995). https://doi.org/10.1007/BF00173193
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00173193