Abstract
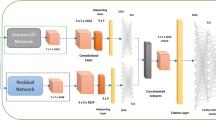
In this paper, we developed cloud-based skin lesion diagnosis system using convolutional neural networks, which consists of the following: (a) Deep learning based classifier that processes user submitted lesion images which runs on a server connected to the cloud based database. (b) Deep learning based classifier performs quality checks and filters user requests before the request is sent off to the diagnosis classifier. (c) A mobile application that runs on Android and iOS platforms to showcase the system. We designed and implemented the system’s architecture.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Rogers, H.W., Weinstock, M.A., Feldman, S.R., Coldiron, B.M.: Incidence estimate of nonmelanoma skin cancer (keratinocyte carcinomas) in the US population, 2012. JAMA Dermatol. 151(10), 1081–1086 (2015)
Cancer Facts and Figures 2018. American Cancer Society. https://www.cancer.org/content/dam/cancer-org/research/cancer-facts-and-statistics/annual-cancer-facts-and-figures/2018/cancer-facts-and-figures-2018.pdf. Accessed 3 May 2018
Stern, R.S.: Prevalence of a history of skin cancer in 2007: results of an incidence-based model. Arch. Dermatol. 146(3), 279–282 (2010)
Guy, G.P., Machlin, S.R., Ekwueme, D.U., Yabroff, K.R.: Prevalence and costs of skin cancer treatment in the U.S., 2002–2006 and 2007–2011. Am. J. Prev. Med. 104(4), e69–e74 (2014). https://doi.org/10.1016/j.amepre.2014.08.036
Siegel, R., Miller, K.D., Jemal, A.: Cancer statistics, 2016. CA Cancer J. Clin. 66, 7–30 (2016)
Kittler, H., Pehamberger, H., Wolf, K., Binder, M.: Diagnostic of dermoscopy. Lancet Oncol. 3, 159–165 (2002)
Yu, L., Chen, H., Dou, Q., Qin, J., Heng, P.A.: Automated melanoma recognition in dermoscopy images via very deep residual networks. IEEE Trans. Med. Imaging 36(4), 994–1004 (2017). https://doi.org/10.1109/TMI.2016.2642839
Esteva, A., Kuprel, B., Novoa, R.A., Ko, J., Swetter, S.M., Blau, H.M., Thrun, S.: Dermatologist-level classification of skin cancer with deep neural networks. Nature 542(7639), 115–118 (2017)
Szegedy, C., Vanhoucke, V., Ioffe, S., Shlens, J., Wojna, Z.: Rethinking the inception architecture for computer vision (2015). Preprint at https://arxiv.org/abs/1512.00567
Russakovsky, O., et al.: Imagenet large scale visual recognition challenge. Int. J. Comput. Vis. 115, 211–252 (2015)
Pan, S.J., Yang, Q.: A survey on transfer learning. IEEE Trans. Knowl. Data Eng. 22, 1345–1359 (2010)
Tschandl, P., Rosendahl, C., Kittler, H.: The HAM10000 dataset, a large collection of multi-source dermatoscopic images of common pigmented skin lesions. Sci. Data 5, 180161 (2018). https://doi.org/10.1038/sdata.2018.161
Codella, N.C.F., Gutman, D., Celebi, M.E., Helba, B., Marchetti, M.A., Dusza, S.W., Kalloo, A., Liopyris, K., Mishra, N., Kittler, H., Halpern, A.: Skin lesion analysis toward Melanoma detection: a challenge. In: 2017 International Symposium on Biomedical Imaging (ISBI), Hosted by the International Skin Imaging Collaboration (ISIC) (2017). arXiv:1710.05006
Cloud Firestore.: (n.d.). https://firebase.google.com/docs/firestore/. Accessed 29 Aug 2018
Chollet, F., and others: Keras, GitHub repository (2018). https://github.com/keras-team/keras
Fei-Fei, L., Fergus, R., Perona, P.: Learning generative visual models from few training examples: an incremental Bayesian approach tested on 101 object categories. IEEE. CVPR 2004, Workshop on Generative-Model Based Vision (2004)
Deng, L.: A tutorial survey of architectures, algorithms, and applications for deep learning. APSIPA Trans. Sig. Inf. Process. 3, e2 (2014)
“Documentation for individual models” (2018). https://keras.io/applications. Accessed 1 Aug 2018
He, K., et al.: Deep residual learning for image recognition. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (2016)
Srivastava, N., et al.: Dropout: a simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 15(1), 1929–1958 (2014)
Ioffe, S., Szegedy, C.: Batch normalization: Accelerating deep network training by reducing internal covariate shift. arXiv preprint (2015). arXiv:1502.03167
Drifty, Inc.: Ionic (2016). https://ionicframework.com
Acknowledgments
The authors gratefully acknowledge funding from NSF award No. 1464537, Industry/University Cooperative Research Center, Phase II under NSF 13-542. We are also thankful to Farris Foundation who also provided funds for this project.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this paper
Cite this paper
Akar, E., Marques, O., Andrews, W.A., Furht, B. (2019). Cloud-Based Skin Lesion Diagnosis System Using Convolutional Neural Networks. In: Arai, K., Bhatia, R., Kapoor, S. (eds) Intelligent Computing. CompCom 2019. Advances in Intelligent Systems and Computing, vol 997. Springer, Cham. https://doi.org/10.1007/978-3-030-22871-2_70
Download citation
DOI: https://doi.org/10.1007/978-3-030-22871-2_70
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-22870-5
Online ISBN: 978-3-030-22871-2
eBook Packages: Intelligent Technologies and RoboticsIntelligent Technologies and Robotics (R0)