Abstract
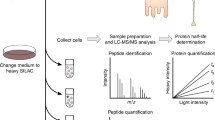
Proteome solubility contains latent information on the nature of protein interaction networks in cells and changes in solubility can provide information on rewiring of networks. Here, we report a simple one-step ultracentrifugation method to separate the soluble and insoluble fraction of the proteome. The method involves quantitative proteomics and a bioinformatics strategy to analyze the changes that arise. Because protein solubility changes are also associated with protein misfolding and aggregation in neurodegenerative disease, we also include a protocol for isolating disease-associated protein aggregates with pulse shape analysis (PulSA) by flow cytometry as a complementary approach that can be used alongside the more general measure of solubility or as a stand-alone approach.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Banani SF, Lee HO, Hyman AA, Rosen MK (2017) Biomolecular condensates: organizers of cellular biochemistry. Nat Rev Mol Cell Biol 18(5):285–298. https://doi.org/10.1038/nrm.2017.7
Ross CA, Poirier MA (2004) Protein aggregation and neurodegenerative disease. Nat Med 10(Suppl):S10–S17. https://doi.org/10.1038/nm1066
Wallace EW, Kear-Scott JL, Pilipenko EV, Schwartz MH, Laskowski PR, Rojek AE, Katanski CD, Riback JA, Dion MF, Franks AM, Airoldi EM, Pan T, Budnik BA, Drummond DA (2015) Reversible, specific, active aggregates of endogenous proteins assemble upon heat stress. Cell 162(6):1286–1298. https://doi.org/10.1016/j.cell.2015.08.041
Huse M (2009) The t-cell-receptor signaling network. J Cell Sci 122(Pt 9):1269–1273. https://doi.org/10.1242/jcs.042762
Sui X, Pires DEV, Ormsby AR, Cox D, Nie S, Vecchi G, Vendruscolo M, Ascher DB, Reid GE, Hatters DM (2020) Widespread remodeling of proteome solubility in response to different protein homeostasis stresses. Proc Natl Acad Sci U S A 117(5):2422–2431. https://doi.org/10.1073/pnas.1912897117
Radwan M, Lilley JD, Ang CS, Reid GE, Hatters DM (2020) Immiscible inclusion bodies formed by polyglutamine and poly(glycine-alanine) are enriched with distinct proteomes but converge in proteins that are risk factors for disease and involved in protein degradation. PLoS One 15(8):e0233247. https://doi.org/10.1371/journal.pone.0233247
Radwan M, Ang CS, Ormsby AR, Cox D, Daly JC, Reid GE, Hatters DM (2020) Arginine in c9orf72 dipolypeptides mediates promiscuous proteome binding and multiple modes of toxicity. Mol Cell Proteomics 19(4):640–654. https://doi.org/10.1074/mcp.RA119.001888
Cox D, Raeburn C, Sui X, Hatters DM (2020) Protein aggregation in cell biology: an aggregomics perspective of health and disease. Semin Cell Dev Biol 99:40–54. https://doi.org/10.1016/j.semcdb.2018.05.003
Kim YE, Hosp F, Frottin F, Ge H, Mann M, Hayer-Hartl M, Hartl FU (2016) Soluble oligomers of polyq-expanded huntingtin target a multiplicity of key cellular factors. Mol Cell 63(6):951–964. https://doi.org/10.1016/j.molcel.2016.07.022
Ramdzan YM, Polling S, Chia CP, Ng IH, Ormsby AR, Croft NP, Purcell AW, Bogoyevitch MA, Ng DC, Gleeson PA, Hatters DM (2012) Tracking protein aggregation and mislocalization in cells with flow cytometry. Nat Methods 9(5):467–470. https://doi.org/10.1038/nmeth.1930
Fonslow BR, Yates JR III (2012) Proteolytic digestion methods for shotgun proteomics. In: Comprehensive sampling and sample preparation. https://doi.org/10.1016/B978-0-12-381373-2.10082-1
Ramdzan YM, Wood R, Hatters DM (2013) Pulse shape analysis (pulsa) to track protein translocalization in cells by flow cytometry: applications for polyglutamine aggregation. Methods Mol Biol 1017:85–93. https://doi.org/10.1007/978-1-62703-438-8_6
Tyanova S, Temu T, Cox J (2016) The maxquant computational platform for mass spectrometry-based shotgun proteomics. Nat Protoc 11(12):2301–2319. https://doi.org/10.1038/nprot.2016.136
Mi H, Muruganujan A, Casagrande JT, Thomas PD (2013) Large-scale gene function analysis with the panther classification system. Nat Protoc 8(8):1551–1566. https://doi.org/10.1038/nprot.2013.092
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13(11):2498–2504. https://doi.org/10.1101/gr.1239303
Tsien RY (1998) The green fluorescent protein. Annu Rev Biochem 67:509–544. https://doi.org/10.1146/annurev.biochem.67.1.509
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Sui, X., Radwan, M., Cox, D., Hatters, D.M. (2022). Probing Protein Solubility Patterns with Proteomics for Insight into Network Dynamics. In: Matějů, D., Chao, J.A. (eds) The Integrated Stress Response. Methods in Molecular Biology, vol 2428. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1975-9_16
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1975-9_16
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1974-2
Online ISBN: 978-1-0716-1975-9
eBook Packages: Springer Protocols