Abstract
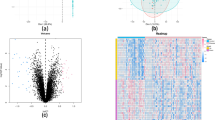
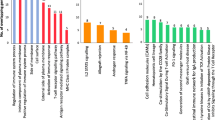
Given that ankylosing spondylitis (AS) occurs in approximately 5 out of 1,000 adults of European descent and the unclear pathogenesis, the aim of the research was to further predict the molecular mechanism of this disease. The Affymetrix chip data GSE25101 were available from Gene Expression Omnibus database. First of all, differentially expressed genes (DEGs) were identified by Limma package in R. Moreover, DAVID was used to perform gene set enrichment analysis of DEGs. In addition, miRanda, miRDB, miRWalk, RNA22 and TargetScan were applied to predict microRNA-target associations. Meanwhile, STRING 9.0 was utilized to collect protein–protein interactions (PPIs) with confidence score >0.4. Then, the PPI networks for up- and down-regulated genes were constructed, and the clustering analysis was undergone using ClusterONE. Finally, protein-domain enrichment analysis of modules was conducted using DAVID. Total 145 DEGs were identified, including 103 up-regulated and 42 down-regulated genes. These DEGs were significantly enriched in phosphorylation (p = 1.21E−05) and positive regulation of gene expression (p = 1.25E−03). Furthermore, one module was screened out from the up-regulated network, which contained 39 nodes and 205 edges. Moreover, the nodes in the module were significantly enriched in ribosomal protein (RPL17, ribosomal protein L17 and MRPL22, mitochondrial ribosomal protein L22) and proteasome (PSMA6, proteasome subunit, alpha type 6, PSMA4)-related domains. Our findings that might explore the potential pathogenesis of AS and RPL17, MRPL22, PSMA6 and PSMA4 have the potential to be the biomarkers for the disease.


Similar content being viewed by others
References
Braun J, Sieper J (2007) Ankylosing spondylitis. Lancet 369(9570):1379–1390
Reveille JD, Sims A-M, Danoy P, Evans DM, Leo P, Pointon JJ, Jin R, Zhou X, Bradbury LA, Appleton LH (2010) Genome-wide association study of ankylosing spondylitis identifies non-MHC susceptibility loci. Nat Genet 42(2):123
Machado P, Landewé R, Lie E, Kvien TK, Braun J, Baker D, van der Heijde D (2011) Ankylosing spondylitis disease activity score (ASDAS): defining cut-off values for disease activity states and improvement scores. Ann Rheum Dis 70(1):47–53
Yekta S, I-h Shih, Bartel DP (2004) MicroRNA-directed cleavage of HOXB8 mRNA. Science 304(5670):594–596
Chatzikyriakidou A, Voulgari P, Georgiou I, Drosos A (2010) The role of microRNA-146a (miR-146a) and its target IL-1R-associated kinase (IRAK1) in psoriatic arthritis susceptibility. Scand J Immunol 71(5):382–385
Thomas GP, Brown MA (2010) Genetics and genomics of ankylosing spondylitis. Immunol Rev 233(1):162–180
Taurog JD (2009) Animal models of spondyloarthritis. Molecular mechanisms of spondyloarthropathies. Springer, Heidelberg, pp 245–254
Díaz-Peña R, Aransay AM, Bruges-Armas J, López-Vázquez A, Rodríguez-Ezpeleta N, Mendibil I, Sánchez A, Torre-Alonso JC, Bettencourt BF, Mulero J (2011) Fine mapping of a major histocompatibility complex in ankylosing spondylitis: association of the HLA-DPA1 and HLA-DPB1 regions. Arthritis Rheum 63(11):3305–3312
Haroon N, Tsui FW, Uchanska-Ziegler B, Ziegler A, Inman RD (2012) Endoplasmic reticulum aminopeptidase 1 (ERAP1) exhibits functionally significant interaction with HLA-B27 and relates to subtype specificity in ankylosing spondylitis. Ann Rheum Dis 71(4):589–595
Barrett T, Troup DB, Wilhite SE, Ledoux P, Rudnev D, Evangelista C, Kim IF, Soboleva A, Tomashevsky M, Edgar R (2007) NCBI GEO: mining tens of millions of expression profiles—database and tools update. Nucleic Acids Res 35(Suppl 1):D760–D765
Pimentel-Santos FM, Ligeiro D, Matos M, Mourão AF, Costa J, Santos H, Barcelos A, Godinho F, Pinto P, Cruz M (2011) Whole blood transcriptional profiling in ankylosing spondylitis identifies novel candidate genes that might contribute to the inflammatory and tissue-destructive disease aspects. Arthritis Res Ther 13(2):R57
Diboun I, Wernisch L, Orengo C, Koltzenburg M (2006) Microarray analysis after RNA amplification can detect pronounced differences in gene expression using limma. BMC Genom 7(1):252
Efron B, Tibshirani R (2002) Empirical Bayes methods and false discovery rates for microarrays. Genet Epidemiol 23(1):70–86
Alvord G, Roayaei J, Stephens R, Baseler MW, Lane HC, Lempicki RA (2007) The DAVID Gene Functional Classification Tool: a novel biological module-centric algorithm to functionally analyze large gene lists. Genome Biol 8:R183
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT (2000) Gene ontology: tool for the unification of biology. Nat Genet 25(1):25–29
John B, Enright AJ, Aravin A, Tuschl T, Sander C, Marks DS (2004) Human microRNA targets. PLoS Biol 2(11):e363
Wang X (2008) miRDB: a microRNA target prediction and functional annotation database with a wiki interface. RNA 14(6):1012–1017
Dweep H, Sticht C, Pandey P, Gretz N (2011) miRWalk–database: prediction of possible miRNA binding sites by “walking” the genes of three genomes. J Biomed Inform 44(5):839–847
Loher P, Rigoutsos I (2012) Interactive exploration of RNA22 microRNA target predictions. Bioinformatics 28(24):3322–3323
Lewis BP, Burge CB, Bartel DP (2005) Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120(1):15–20
Von Mering C, Huynen M, Jaeggi D, Schmidt S, Bork P, Snel B (2003) STRING: a database of predicted functional associations between proteins. Nucleic Acids Res 31(1):258–261
Smoot ME, Ono K, Ruscheinski J, Wang P-L, Ideker T (2011) Cytoscape 2.8: new features for data integration and network visualization. Bioinformatics 27(3):431–432
Nepusz T, Yu H, Paccanaro A (2012) Detecting overlapping protein complexes in protein–protein interaction networks. Nat Methods 9(5):471–472
Apweiler R, Attwood TK, Bairoch A, Bateman A, Birney E, Biswas M, Bucher P, Cerutti L, Corpet F, Croning MDR (2001) The InterPro database, an integrated documentation resource for protein families, domains and functional sites. Nucleic Acids Res 29(1):37–40
Anderson JJ, Baron G, Van Der Heijde D, Felson DT, Dougados M (2001) Ankylosing spondylitis assessment group preliminary definition of short-term improvement in ankylosing spondylitis. Arthritis Rheum 44(8):1876–1886
Zai-Xing Y, Yan L, Hao W, Ye Z, Chang L, Ren-Qian Z (2008) Preliminary clinical measurement of the expression of TNF-related apoptosis inducing ligand in patients with ankylosing spondylitis. J Clin Lab Anal 22(2):138–145
Ciccia F, Accardo-Palumbo A, Giardina A, Di Maggio P, Principato A, Bombardieri M, Rizzo A, Alessandro R, Ferrante A, Principe S (2010) Expansion of intestinal CD4+ CD25 high Treg cells in patients with ankylosing spondylitis: a putative role for interleukin-10 in preventing intestinal Th17 response. Arthritis Rheum 62(12):3625–3634
Shaw J, Kollnberger S (2012) New perspectives on the ligands and function of the killer cell immunoglobulin-like receptor KIR3DL2 in health and disease. Front Immunol 3:339
DiMauro S, Schon EA (2008) Mitochondrial disorders in the nervous system. Annu Rev Neurosci 31:91–123
Wendling D, Didier J, Vuitton D (1991) The phagocyte oxidative metabolism function in ankylosing spondylitis. Rheumatol Int 11(4–5):187–189
Smolock EM, Korshunov VA, Glazko G, Qiu X, Gerloff J, Berk BC (2012) Ribosomal protein L17, RpL17, is an inhibitor of vascular smooth muscle growth and carotid intima formation. Circulation 126(20):2418–2427
Cmejla R, Cmejlova J, Handrkova H, Petrak J, Pospisilova D (2007) Ribosomal protein S17 gene (RPS17) is mutated in Diamond–Blackfan anemia. Hum Mutat 28(12):1178–1182
Mees ST, Mardin WA, Wendel C, Baeumer N, Willscher E, Senninger N, Schleicher C, Colombo-Benkmann M, Haier J (2010) EP300—A miRNA-regulated metastasis suppressor gene in ductal adenocarcinomas of the pancreas. Int J Cancer 126(1):114–124
Yu F, Cui Y, Zhou X, Zhang X, Han J (2011) Osteogenic differentiation of human ligament fibroblasts induced by conditioned medium of osteoclast-like cells. Biosci Trends 5(2):46
Graham LM, Gupta V, Schafer G, Reid DM, Kimberg M, Dennehy KM, Hornsell WG, Guler R, Campanero-Rhodes MA, Palma AS (2012) The C-type lectin receptor CLECSF8 (CLEC4D) is expressed by myeloid cells and triggers cellular activation through Syk kinase. J Biol Chem 287(31):25964–25974
Cismasiu VB, Ghanta S, Duque J, Albu DI, Chen H-M, Kasturi R, Avram D (2006) BCL11B participates in the activation of IL2 gene expression in CD4+ T lymphocytes. Blood 108(8):2695–2702
Wright C, Edelmann M, Kollnberger S, Kramer H, McGowan S, McHugh K, Taylor S, Kessler B, Bowness P (2009) Ankylosing spondylitis monocytes show upregulation of proteins involved in inflammation and the ubiquitin proteasome pathway. Ann Rheum Dis 68(10):1626–1632
Aldea G, Výprachtický D, Cimrová V (2004) Modification of poly (styrene-alt-maleic anhydride) with 1,3,4-oxadiazole units for electroluminescent devices. Macromolecular symposia. Wiley, Hoboken, pp 523–528
Haroon N, Maksymowych WP, Rahman P, Tsui FW, O’Shea FD, Inman RD (2012) Radiographic severity of ankylosing spondylitis is associated with polymorphism of the large multifunctional peptidase 2 gene in the spondyloarthritis research consortium of Canada cohort. Arthritis Rheum 64(4):1119–1126
van Endert P (2011) Post-proteasomal and proteasome-independent generation of MHC class I ligands. Cell Mol Life Sci 68(9):1553–1567
Parkes M, Cortes A, van Heel DA, Brown MA (2013) Genetic insights into common pathways and complex relationships among immune-mediated diseases. Nat Rev Genet 14(9):661–673
Acknowledgments
This work was supported by Shanghai Municipal Health and Family Planning Commission (Project No. 2012QL044A).
Conflict of interest
The authors have declared that no competing interests exist.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhao, H., Wang, D., Fu, D. et al. Predicting the potential ankylosing spondylitis-related genes utilizing bioinformatics approaches. Rheumatol Int 35, 973–979 (2015). https://doi.org/10.1007/s00296-014-3178-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00296-014-3178-9